---
title: "Lecture 08"
subtitle: "Generalized Measurement Models: Modeling Observed Data II"
author: "Jihong Zhang"
institute: "Educational Statistics and Research Methods"
title-slide-attributes:
data-background-image: ../Images/title_image.png
data-background-size: contain
execute:
echo: true
eval: false
format:
html:
code-tools: true
code-line-numbers: false
code-fold: show
code-summary: 'Click here to see code'
number-sections: true
number-offset: 0
fig-width: 7
fig-align: center
message: false
revealjs:
output-file: slides-index.html
logo: ../Images/UA_Logo_Horizontal.png
incremental: false # choose "false "if want to show all together
transition: slide
background-transition: fade
theme: [simple, ../pp.scss]
footer: <https://jihongzhang.org/teaching/2024-01-12-syllabus-adv-multivariate-esrm-6553>
scrollable: true
slide-number: true
chalkboard: true
number-sections: false
code-line-numbers: true
code-annotations: below
code-copy: true
code-summary: ''
highlight-style: arrow
view: 'scroll' # Activate the scroll view
scrollProgress: true # Force the scrollbar to remain visible
mermaid:
theme: neutral
#bibliography: references.bib
---
# Review Previous Class
## In Previous Class...
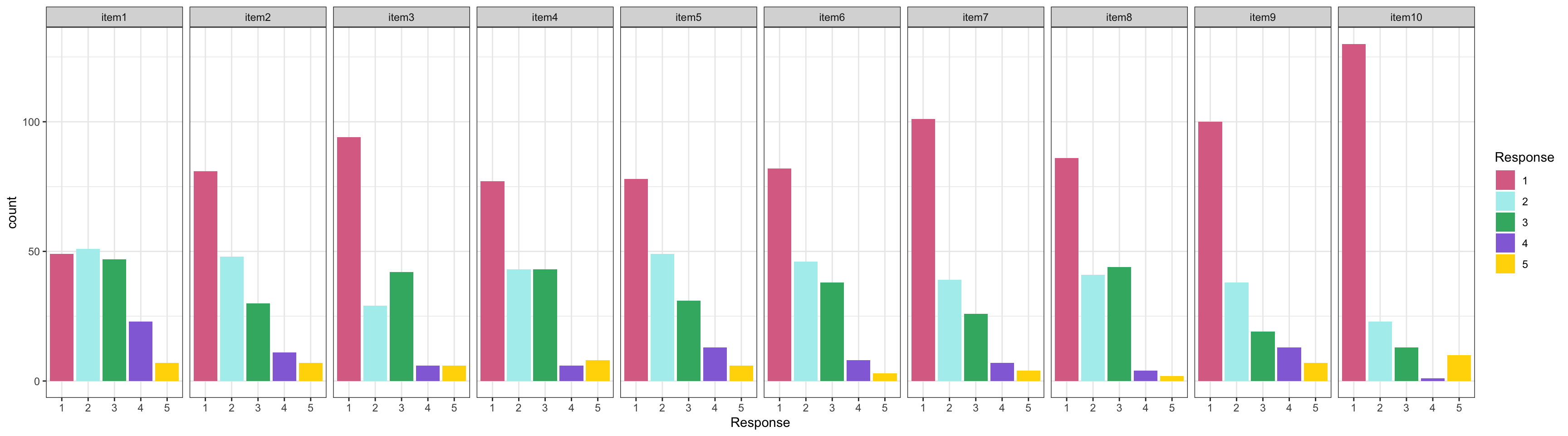
1. We introduced a self-reported survey data, called `Conspiracy Theories`
- The scale has 10 items with 5-point Likert scale response
- Items have varied item difficult and Positive skewed item response distribution
2. We talked about using if factor analysis is a proper method when normality assumption is violated
3. We checked the item characteristics curves
```{r}
#| fig-width: 8
#| eval: true
library(tidyverse)
library(here)
library(blavaan)
self_color <- c("#DB7093", "#AFEEEE", "#3CB371", "#9370DB", "#FFD700")
root_dir <- "teaching/2024-01-12-syllabus-adv-multivariate-esrm-6553/Lecture07/Code"
dat <- read.csv(here(root_dir, 'conspiracies.csv'))
itemResp <- dat[,1:10]
colnames(itemResp) <- paste0('item', 1:10)
conspiracyItems = itemResp
itemResp |>
rownames_to_column("ID") |>
pivot_longer(-ID, names_to = "Item", values_to = "Response") |>
mutate(Item = factor(Item, levels = paste0('item', 1:10)),
Response = factor(Response, levels = 1:5)) |>
ggplot() +
geom_bar(aes(x = Response, fill = Response, group = Response),
position = position_stack()) +
facet_wrap(~ Item, nrow = 1, ncol = 10) +
theme_bw() +
scale_fill_manual(values = self_color)
```
## Today's Lecture Objectives
1. Dive deep into factor scoring
2. Show how different initial values affect Bayesian model estimation
3. Show how parameterization differs for standardized latent variables vs. marker item scale identification
::: {.callout-note appearance="minimal"}
**Lecture 07 Files:** [{{< fa file-code >}} Lecture07.R](../Lecture07/Code/Lecture07.R){download="Lecture07.R"} | [{{< fa file-csv >}} conspiracies.csv](../Lecture07/Code/conspiracies.csv){download="conspiracies.csv"}
:::
## Posterior Distribution for Item Parameters
Before moving onto the latent variable, let's note the posterior distribution of item parameters (for a single item):
$$
f(\mu_i,\lambda_i,\psi_i\mid\boldsymbol{Y})\propto f(\boldsymbol{Y}\mid\mu_i,\lambda_i,\psi_i)f(\mu_i, \lambda_i,\psi_i)
$$
where $f(\boldsymbol{Y}\mid\mu_i,\lambda_i,\psi_i)$ is the (joint) posterior distribution of the parameters for item $i$ conditional on the adta
$f(\boldsymbol{Y}\mid\mu_i,\lambda_i,\psi_i)$ is the distribution we defined for our observed data:
$$
f(\boldsymbol{Y}\mid\mu_i,\lambda_i,\psi_i)\sim N(\mu_i+\lambda_i\theta_p, \psi_i)
$$
$f(\mu_i, \lambda_i, \psi_i)$ is the (joint) prior distribution for each of the parameters, which, are independent:
$$
f(\mu_i, \lambda_i, \psi_i) = f(\mu_i)f(\lambda_i)f(\psi_i)
$$
## Investigating the Latent Variables
The estimated latent variables are then:
```{r}
#| eval: true
save_dir <- "~/Library/CloudStorage/OneDrive-Personal/2024_Spring/ESRM6553 - Advanced Multivariate Modeling/Lecture07"
modelCFA_samples <- readRDS(here(save_dir, "model01.RDS"))
modelCFA_samplesFail <- readRDS(here(save_dir, "model01fail.RDS"))
modelCFA_samplesFix <- readRDS(here(save_dir, "model03fix.RDS"))
model01_blv <- readRDS(here(save_dir, "model01_blv.RDS"))
```
```{r}
#| eval: true
modelCFA_samples$summary('theta')
```
## EAP Estimates of Latent Variables
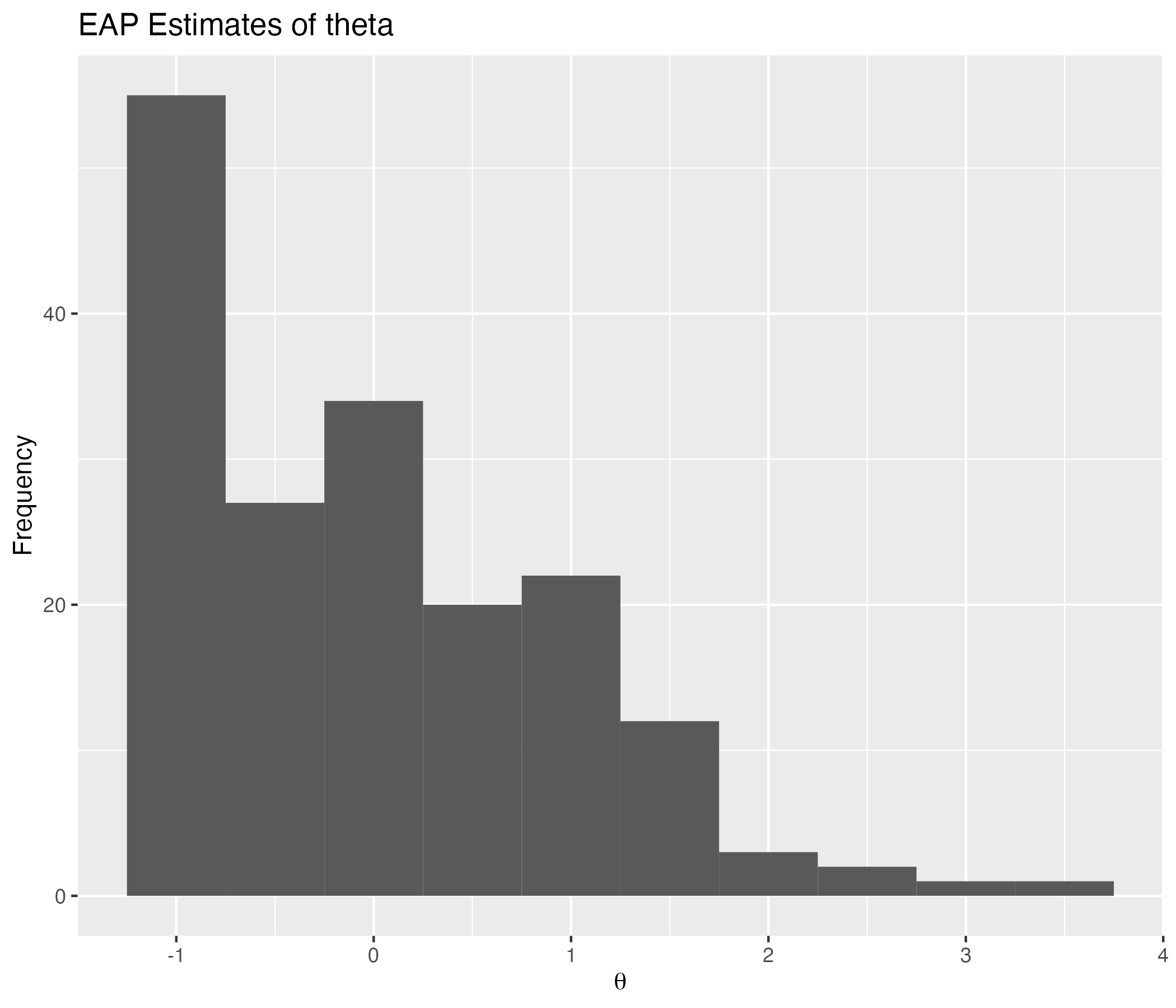{width=70%}
## Density of EAP Estimates
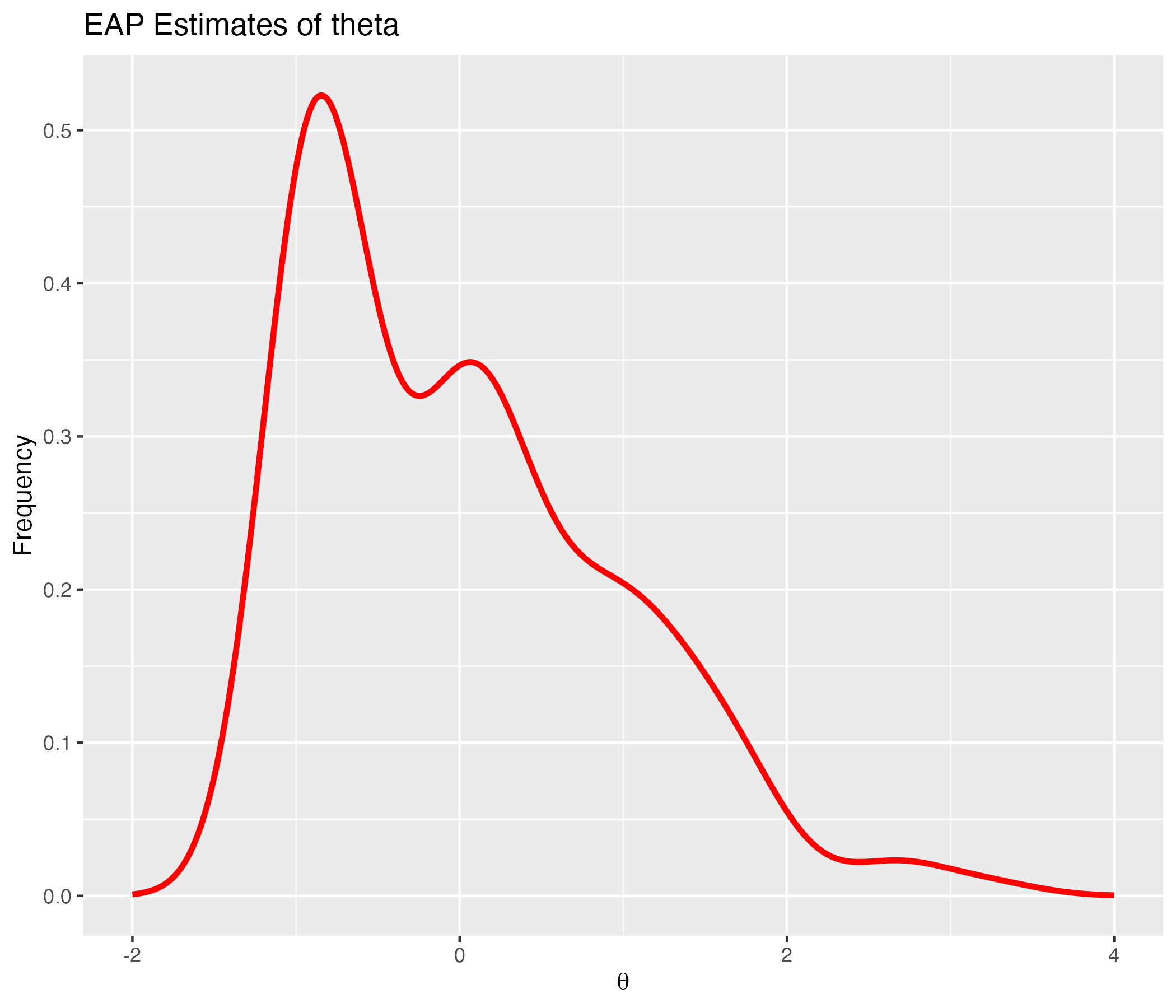{width=70%}
## Density of 500 Posterior draws of θ
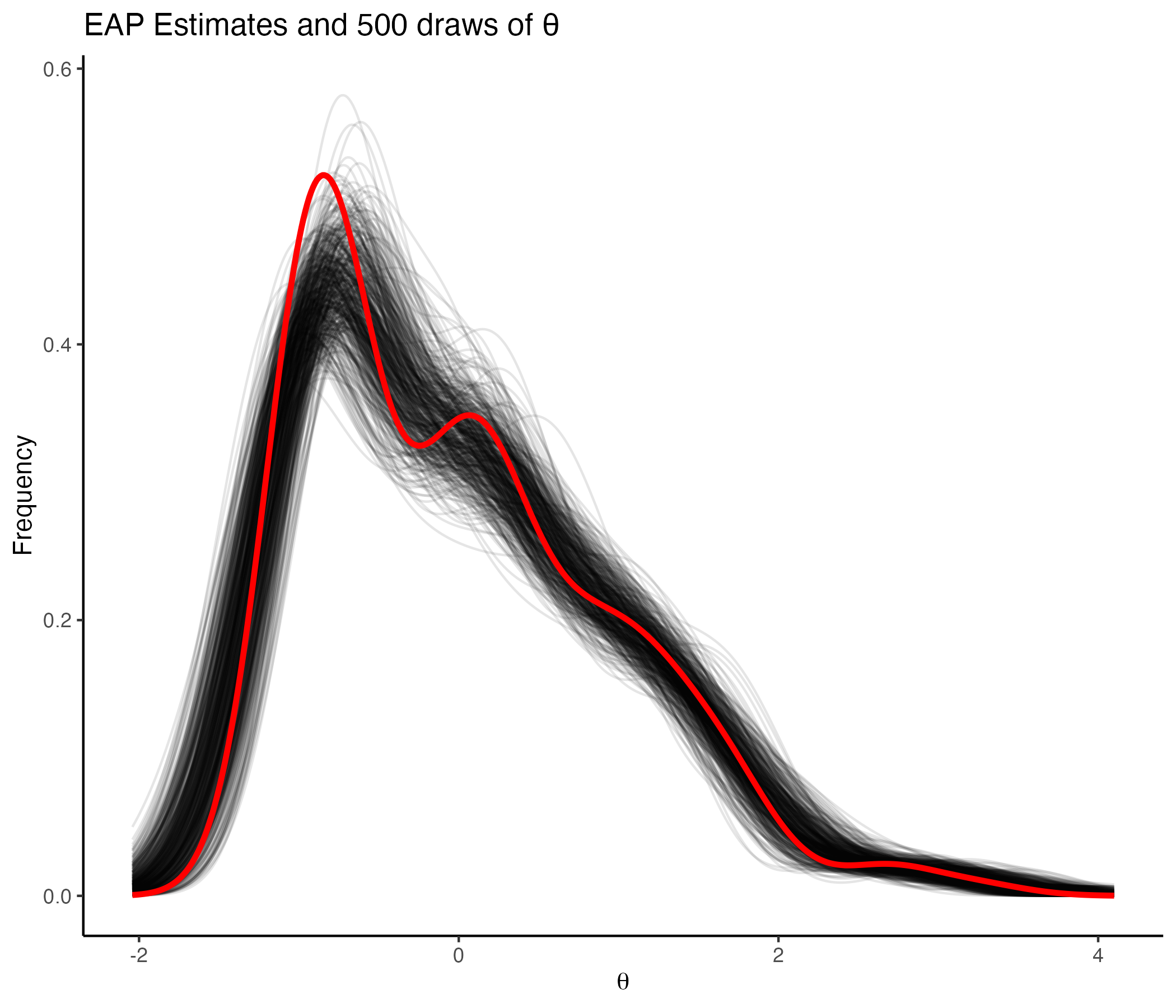{width=70%}
## Comparing Posterior Distribution for Individuals
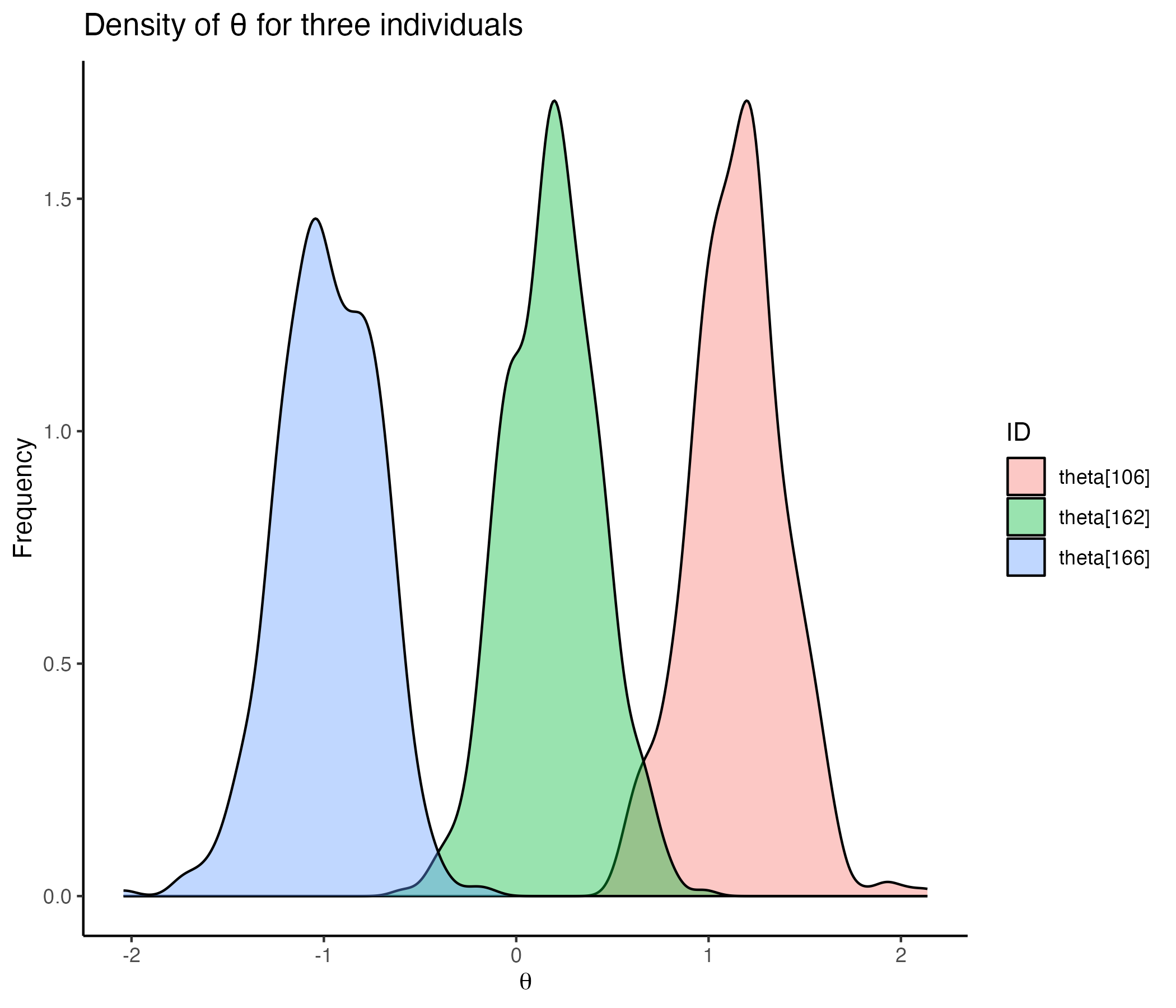{width=70%}
## Comparing EAP Estimate with Posterior SD
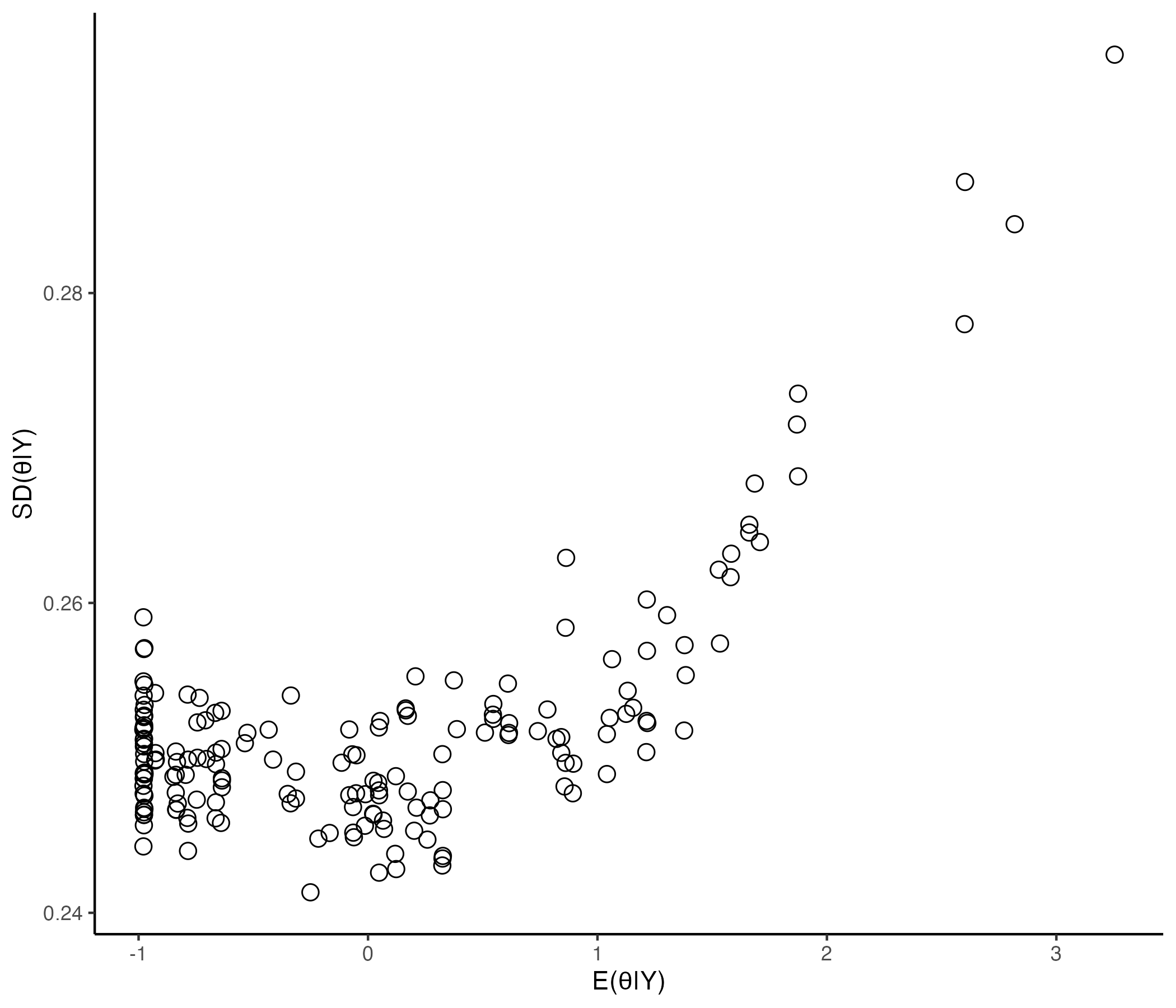{width=70%}
## Comparing EAP Estimate with Sum Score
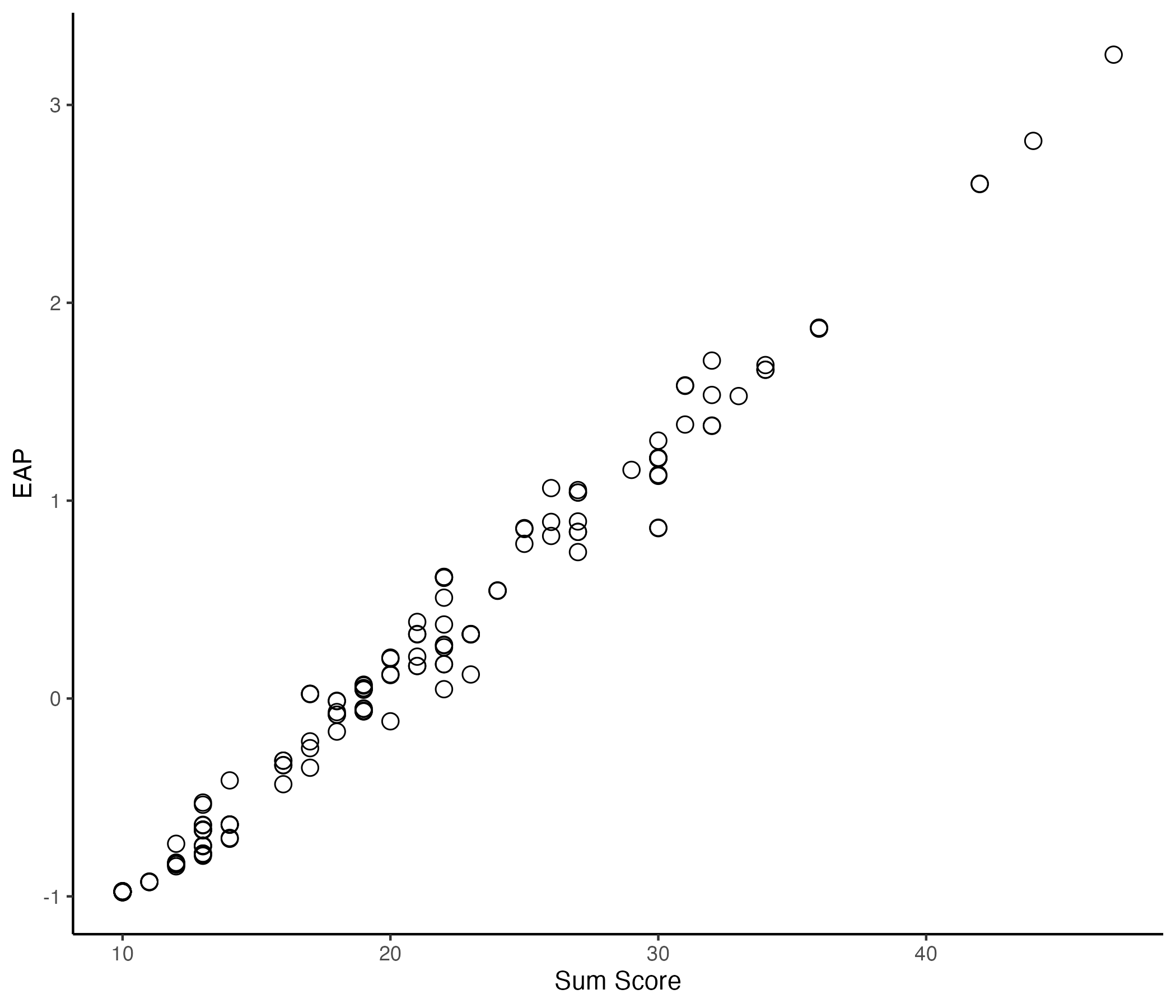{width=70%}
## Comparing EAP Estimate with Factor Score by `lavaan`
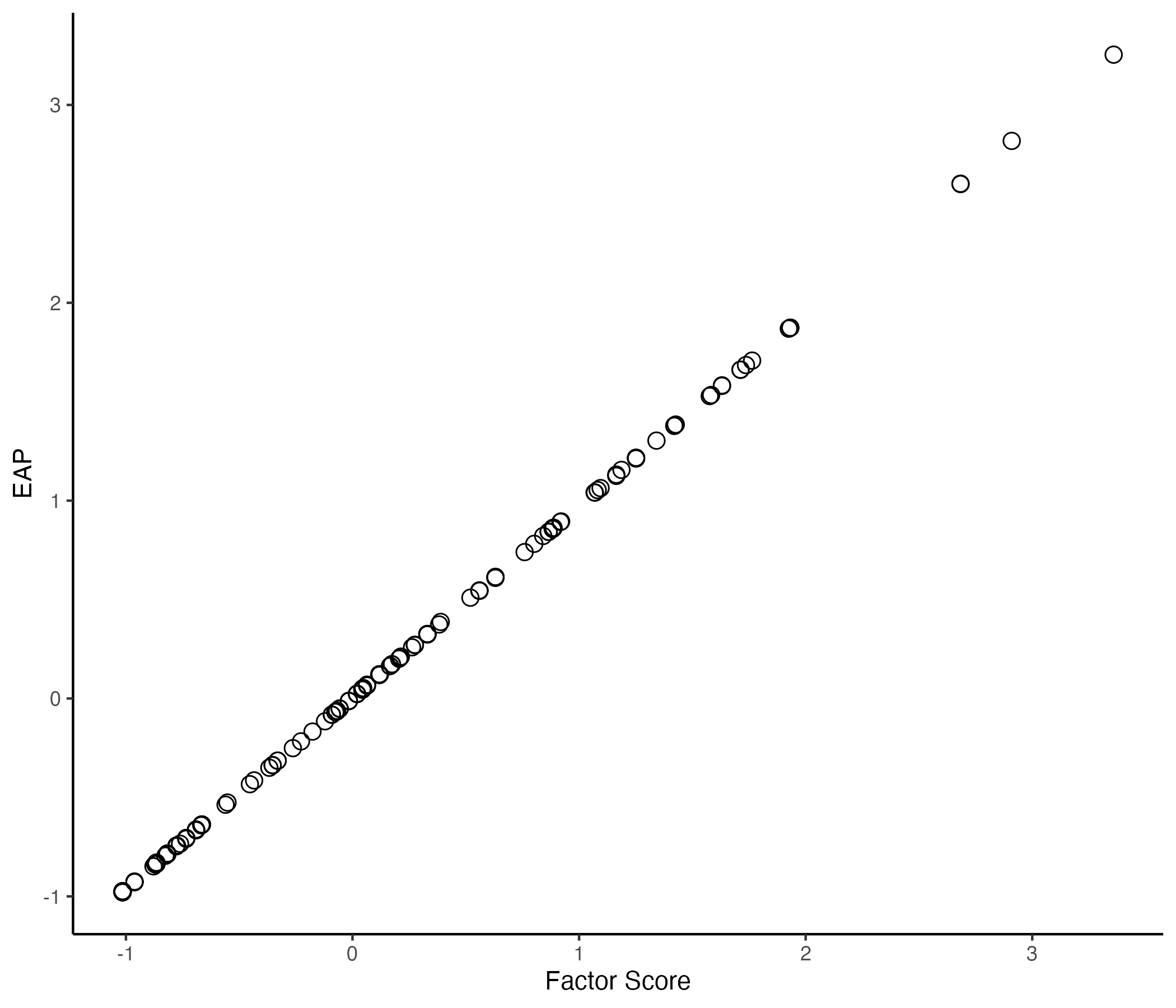{width=70%}
## Posterior Distribution for Person Parameter θ
The posterior distribution of the person parameters (the latent variable for a single person):
$$
f(\theta_p\mid\boldsymbol{Y}) \propto f(\boldsymbol{Y} \mid \theta_p)f(\theta_p)
$$
Here:
- $f(\theta_p\mid\boldsymbol{Y})$ is the posterior distribution of the latent variable conditional on the observed data;
- $f(\boldsymbol{Y} \mid \theta_p)$ is the model (data) likelihood:
$$
f(\boldsymbol{Y} \mid \theta_p) = \prod_{i=1}^{I} f(Y_i\mid\theta_p)
$$
- $f(Y_i \mid \theta_p)$ is one individual item's data likelihood: $f(Y_i \mid \theta_p) \sim N(\mu_i+\lambda_i\theta_p, \psi_i)$;
- $f(\theta_p) \sim N(0,1)$ is the prior distribution for the latent variable – $\theta_p$
# Measurement Model Estimation Fails
## Recall: Stan's `parameters {}` Block
```{stan, output.var='display'}
#| echo: true
parameters {
vector[nObs] theta; // the latent variables (one for each person)
vector[nItems] mu; // the item intercepts (one for each item)
vector[nItems] lambda; // the factor loadings/item discriminations (one for each item)
vector<lower=0>[nItems] psi; // the unique standard deviations (one for each item)
}
```
Here, the parameterization of $\lambda$ (factor loadings / discrimination parameters) can lead to problems in estimation
- The issue is $\lambda_i\theta_p=(-\lambda_i)(-\theta_p)$
- Depending on the random starting values of each of these parameters (per Markov chain), a given chain may converge to different region with others
- To demonstrate, we will start with different random number seend
- Currently use 09102022: works fine
- Change to 25102022: big problem
## New Samples Syntax
```{r}
#| echo: true
#| code-line-numbers: "|3"
modelCFA_samplesFail = modelCFA_stan$sample(
data = modelCFA_data,
seed = 25102022,
chains = 4,
parallel_chains = 4,
iter_warmup = 1000,
iter_sampling = 2000
)
```
Convergence fail with maximum of $\hat R$ as:
```{r}
#| eval: true
max(modelCFA_samplesFail$summary(.cores =4)$'rhat')
```
## Why Convergence Failed
- The issue of exchangeable likelihood: $\lambda_i\theta_p=(-\lambda_i)(-\theta_p)$
```{r}
#| eval: true
#| echo: false
modelCFA_samplesFail$summary('lambda', .cores =4) |> mutate(across(-variable, \(x) round(x, 3)))
```
## Posterior Trace Plots of λ
Unfortunately, we are unable to extract initial values that were generated automatically by `cmdstanr`. See [here](https://mc-stan.org/cmdstanr/reference/fit-method-init.html) for more details.
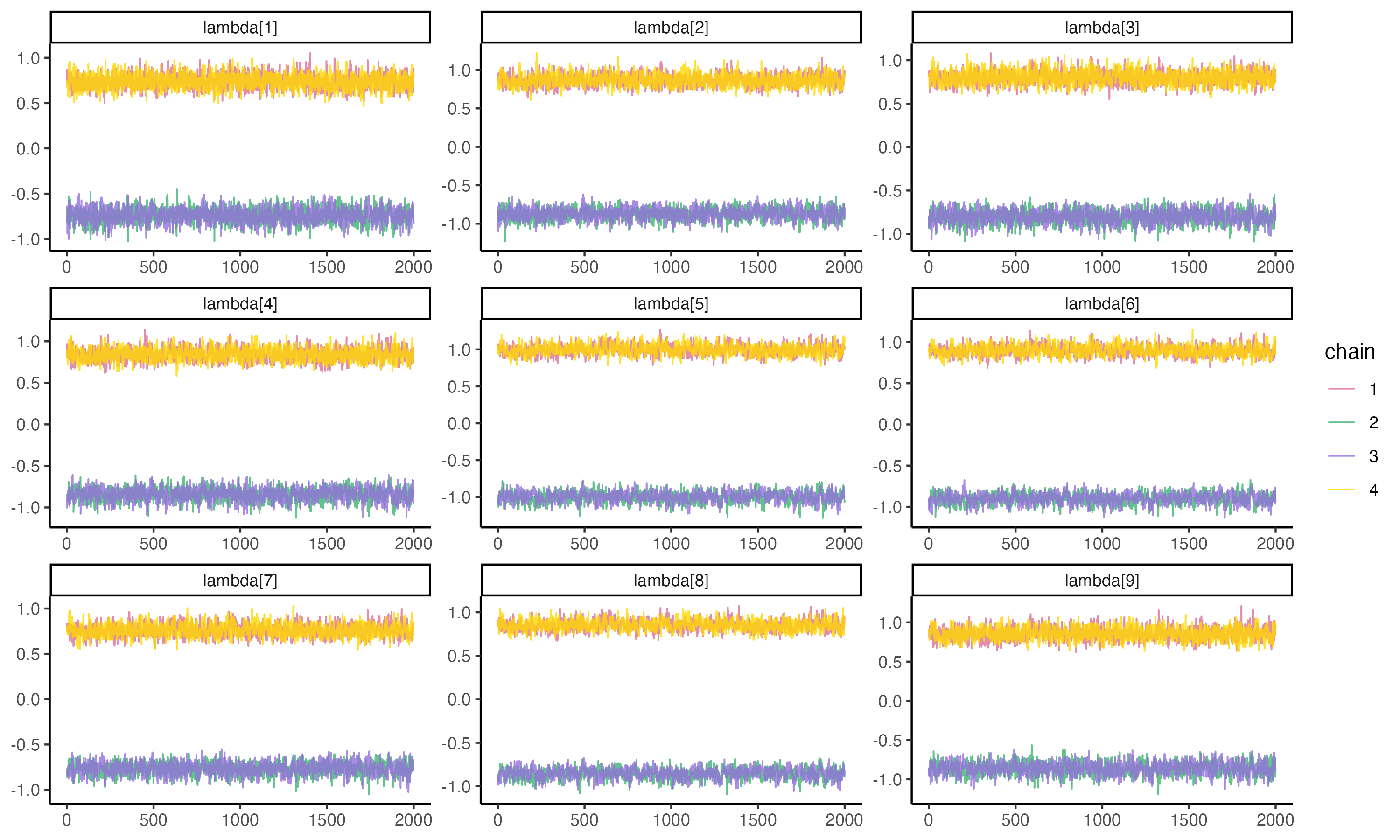{fig-align="center" width=70%}
## Posterior Density Plots of λ
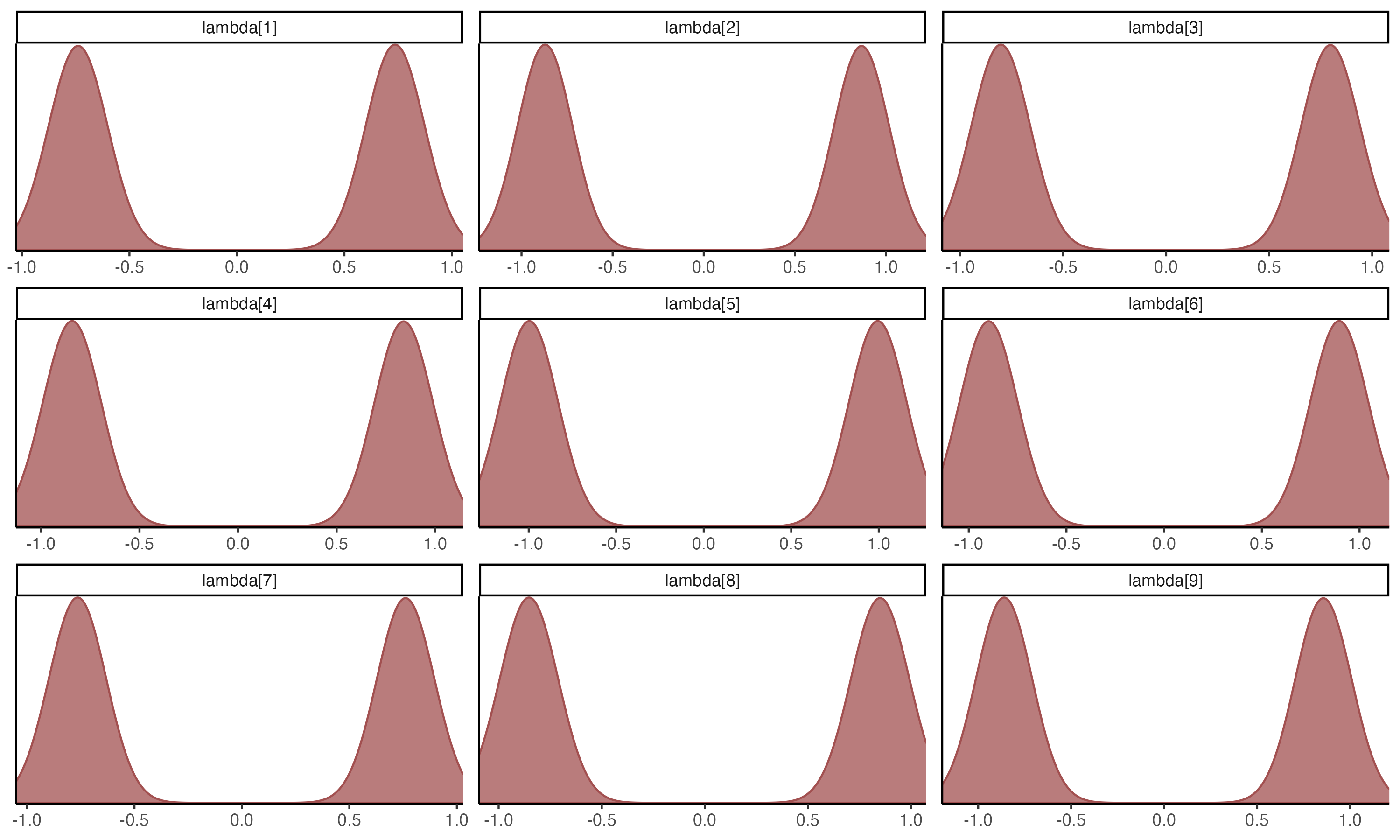{fig-align="center" width=70%}
## Examining Latent Variable
```{r}
#| eval: true
#| echo: false
print(modelCFA_samplesFail$summary('theta', .cores = 4) |> mutate(across(c(mean, median), \(x) round(x, 3))), n = 30)
```
## Posterior Trace Plots of θ
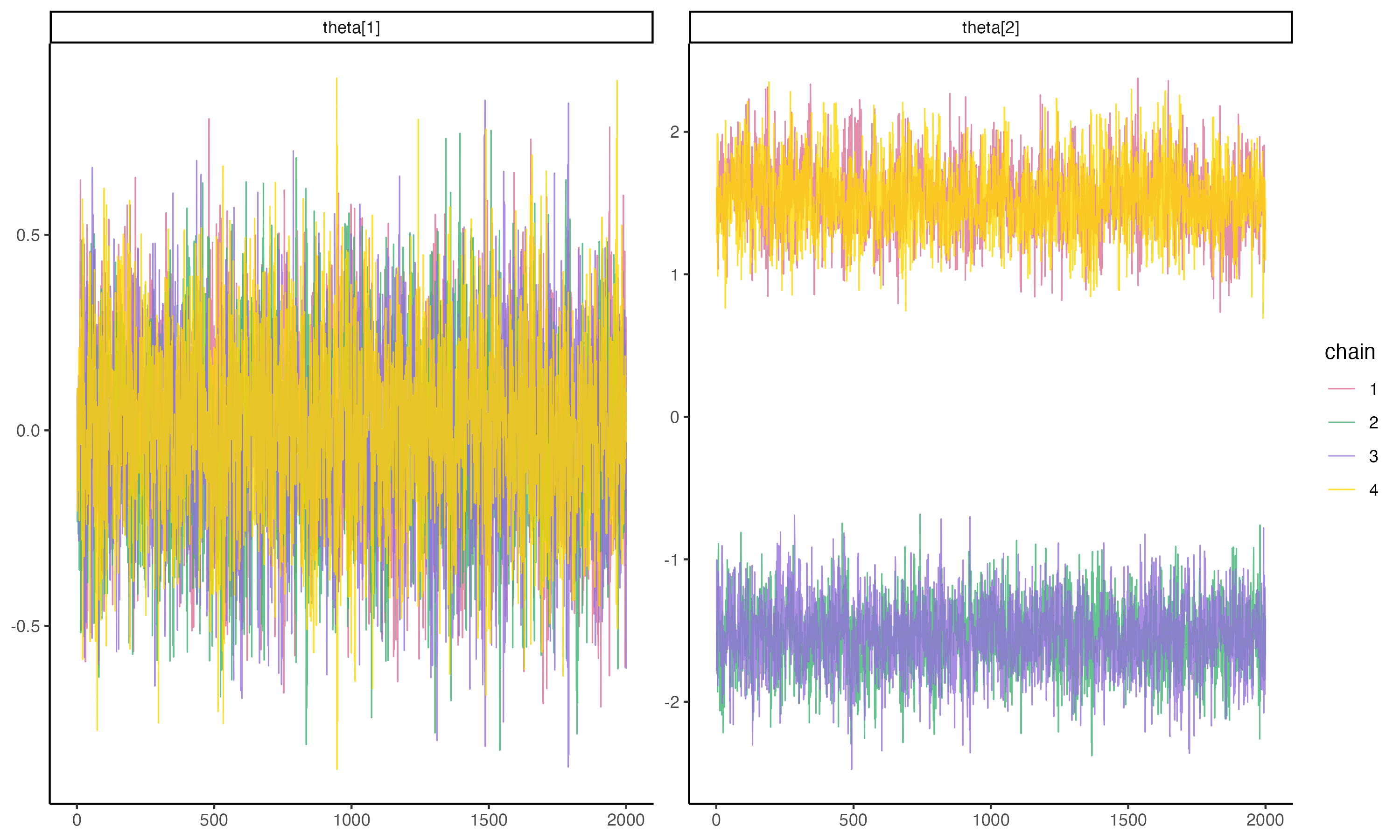{fig-align="center" width=70%}
## Posterior Density Plots of θ
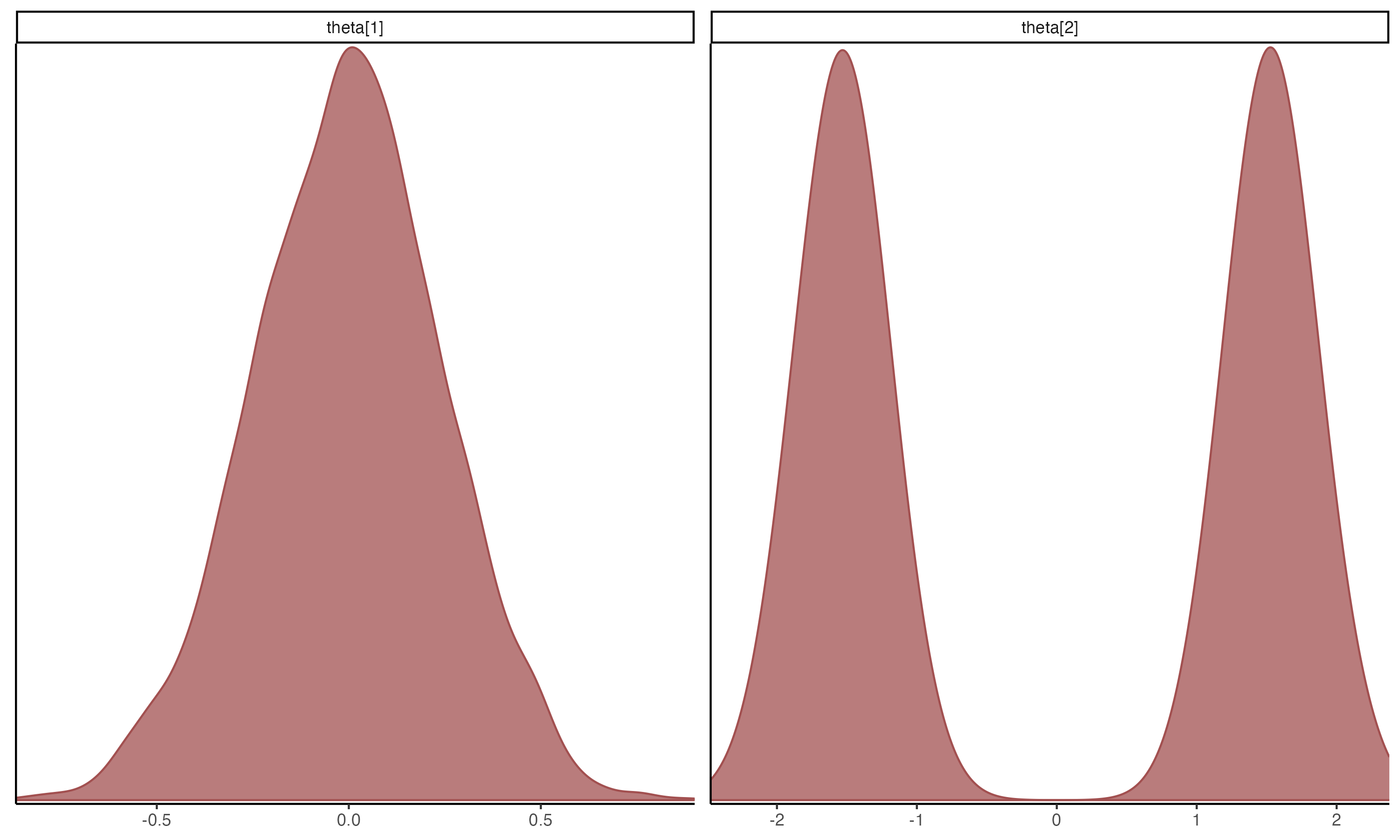{width=70%}
## Fixing Convergence
`Stan` allows starting values to be set via `cmdstanr`
- Documentation is very lacking, I can show you one method but it may change in the future.
Alternatively:
- In `Stan` file, restrict $\lambda$ to be positive: `vector<lower=0>[nItems] lambda;`
- Can also choose prior that has strictly positive range (like log-normal distribution)
- Note: the restriction on the space of $\lambda$ will not permit truely negative values
- Not ideals as negative $\lambda$ values are informative as a problem with data
```{stan, output.var='display'}
#| echo: true
parameters {
vector[nObs] theta; // the latent variables (one for each person)
vector[nItems] mu; // the item intercepts (one for each item)
vector<lower=0>[nItems] lambda; // the factor loadings/item discriminations (one for each item)
vector<lower=0>[nItems] psi; // the unique standard deviations (one for each item)
}
```
## Setting Starting (Inital) Values in `Stan`
Starting values (initial values) are the first values used when an MCMC chain starts
- In `Stan`, by default, parameters are randomly started between -2 and 2
- Bounded parameters are transformed so they are unbounded in the algorithm
- What we need:
- Randomly start all $\lambda$ parameters so that they converge to the $\lambda_i\theta_p$ mode
- As opposed to the (-$\lambda_i$)(-$\theta_p$) mode
## `cmdstanr` Syntax for Initial Values
Add the init option to the `$sample()` function of the `cmdstanr` object:
```{stan, output.var='display'}
#| echo: true
# set starting values for some of the parameters
modelCFA_samples2fixed = modelCFA_stan$sample(
data = modelCFA_data,
seed = 25102022,
chains = 4,
parallel_chains = 4,
iter_warmup = 2000,
iter_sampling = 2000,
init = function() list(lambda=rnorm(nItems, mean=10, sd=2))
)
```
The `init` option can be specified as a function, here, randomly starting each $\lambda$ following a normal distribution
## Initialization Process
See the lecture R syntax for information on how to confirm starting values are set.
You should find the initial values using `$init()` function
```{r}
#| eval: true
#| echo: true
modelCFA_samplesFix$init()
```
## Final Results: Parameters
```{r}
#| eval: true
#| echo: true
# Check convergence
max(modelCFA_samplesFix$summary()$'rhat')
print(modelCFA_samplesFix$summary(c("mu", "lambda", "psi"),.cores = 4) |> mutate(across(c(mean, median), \(x) round(x, 3))), n = Inf)
```
## Comparing Results with different inits
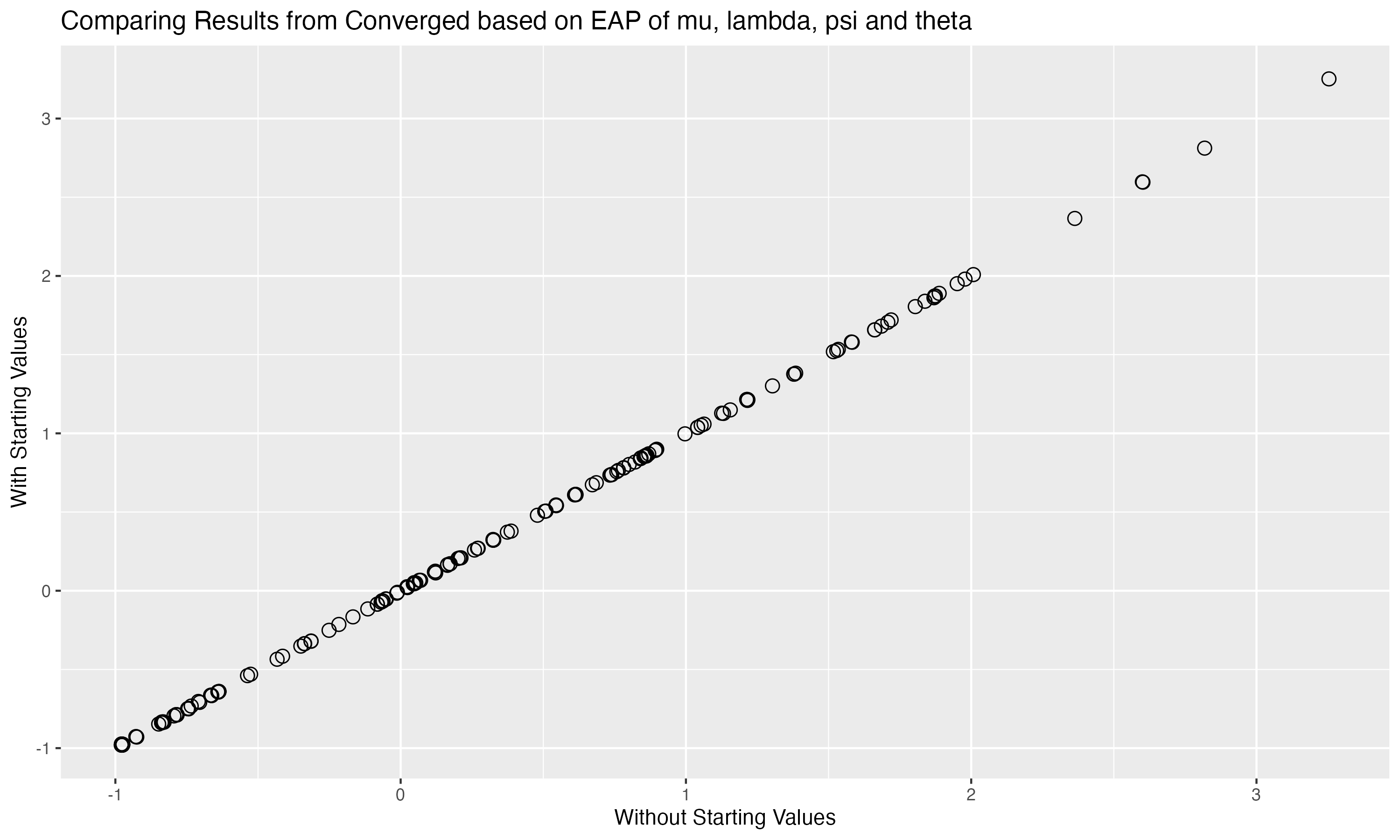{fig-align="center" width=70%}
Correlation of all parameters across both algorithm runs:
```{r}
#| echo: true
#| eval: true
cor(modelCFA_samplesFix$summary(variables = c("mu", "lambda", "psi", "theta"), .cores = 4)$mean,
modelCFA_samples$summary(variables = c("mu", "lambda", "psi", "theta"), .cores = 4)$mean)
```
## CFA model with `blavaan`
```{r}
#| echo: true
blavaan.model <- ' theta =~ item1 + item2 + item3 + item4 + item5 + item6 + item7 + item8 + item9 + item10 '
model01_blv <- bcfa(blavaan.model, data=conspiracyItems,
n.chains = 4, burnin = 1000, sample = 2000,
target = 'stan', seed = 09102022,
save.lvs = TRUE, # save sampled latent variable
std.lv = TRUE,
bcontrol = list(cores = 4),
mcmcfile=TRUE # save Stan file
) # standardized latent variable
```
------------------------------------------------------------------------
### `blavaan` Results
```{r}
#| eval: true
#| echo: true
summary(model01_blv)
```
------------------------------------------------------------------------
### Comparing factors score
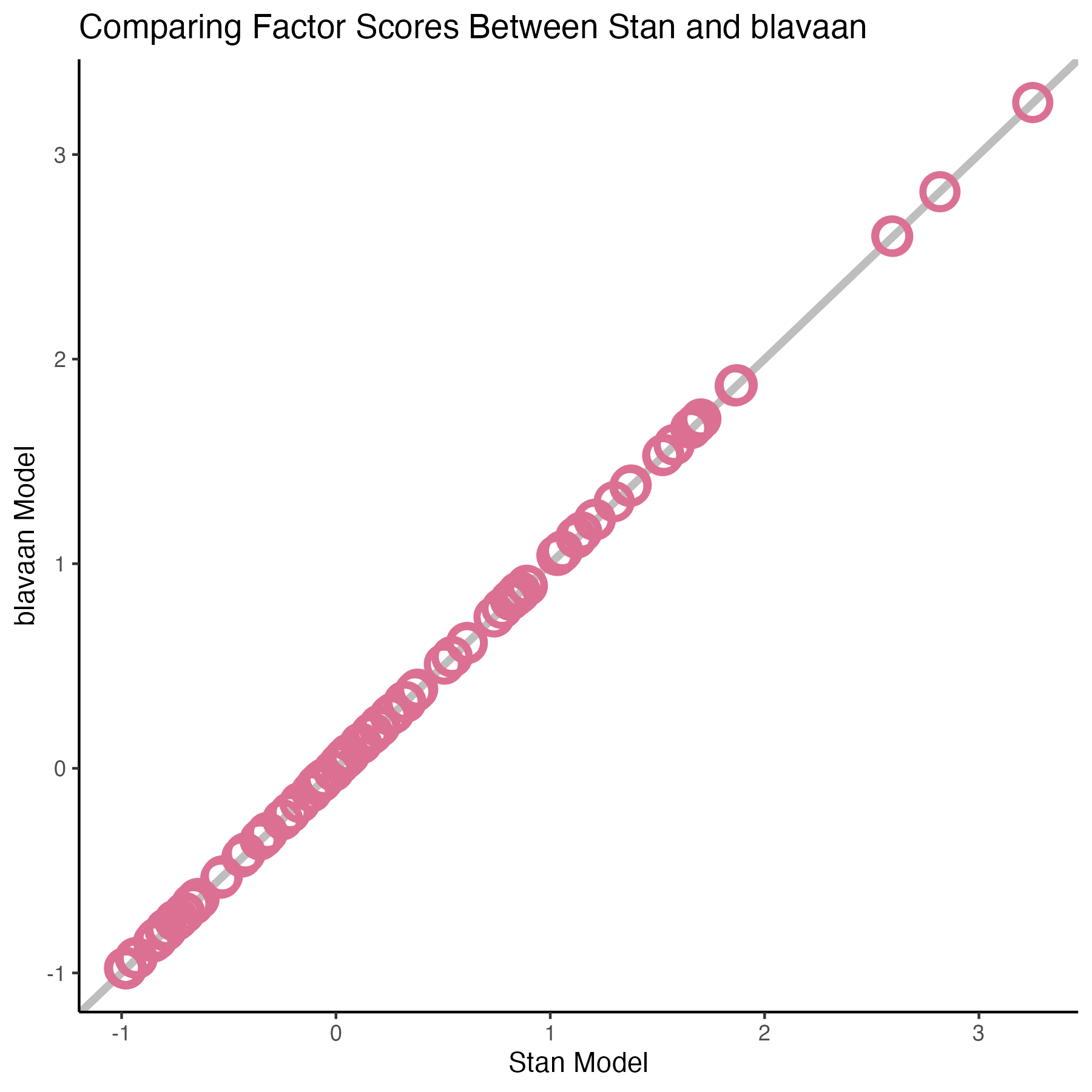{fig-align="center" width=70%}
------------------------------------------------------------------------
### Comparing Parameters
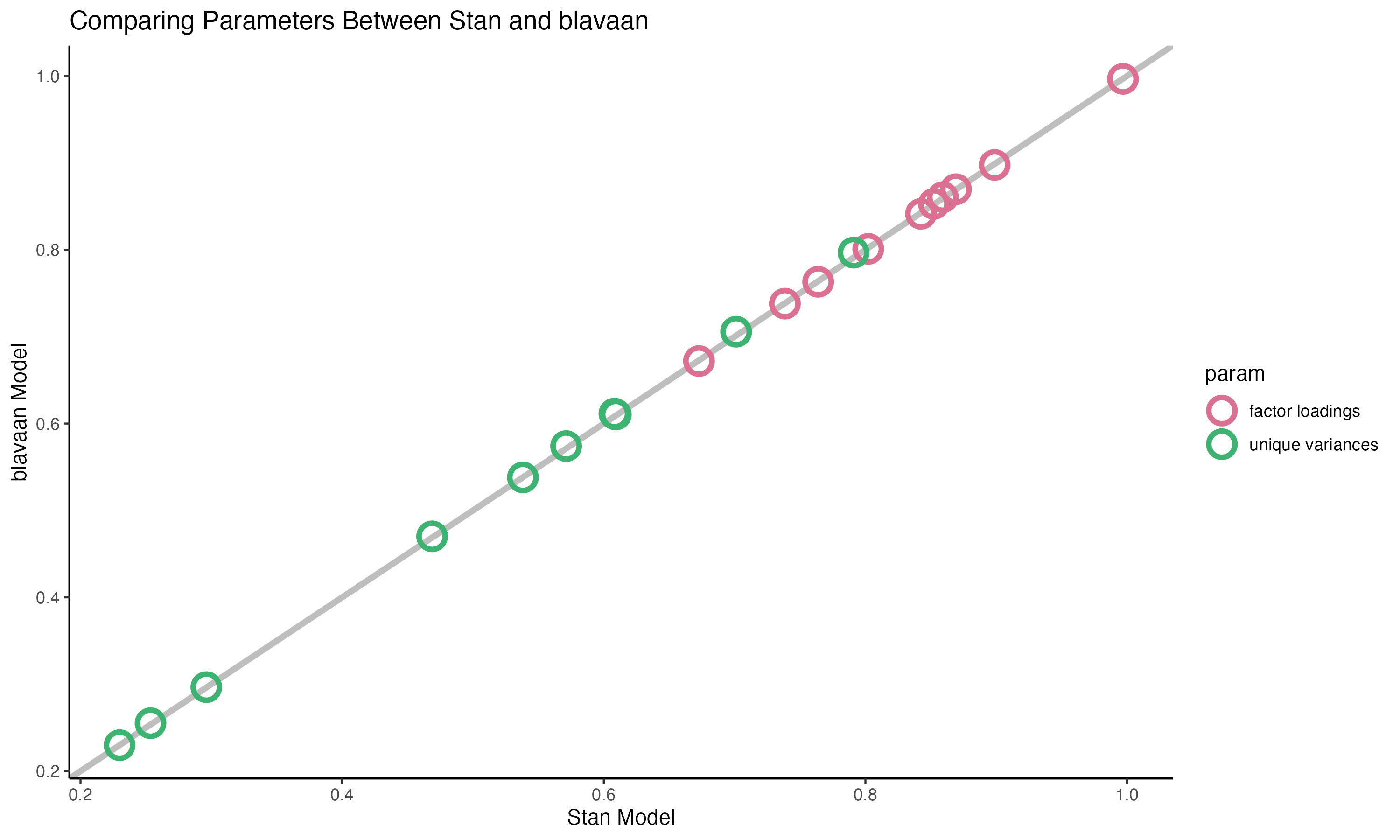{fig-align="center" width=70%}
------------------------------------------------------------------------
### Limitation of blavaan 🧛🏼
You can find more details in the [Edgar Merkle's paper](https://www.jstatsoft.org/article/view/v100i06).
- Very unreadable Stan file (1891 lines)
- limits in setting up prior distributions (cannot change distribution family for lambda and psi)
- Use `rstan` rather than `cmdstanr`
```{stan, output.var='display'}
#| echo: true
/* This file is based on LERSIL.stan by Ben Goodrich.
https://github.com/bgoodri/LERSIL */
functions { // you can use these in R following `rstan::expose_stan_functions("foo.stan")`
// mimics lav_mvnorm_cluster_implied22l():
matrix calc_W_tilde(matrix sigma_w, vector mu_w, array[] int var1_idx, int p_tilde) {
matrix[p_tilde, p_tilde + 1] out = rep_matrix(0, p_tilde, p_tilde + 1); // first column is mean vector
vector[p_tilde] mu1 = rep_vector(0, p_tilde);
matrix[p_tilde, p_tilde] sig1 = rep_matrix(0, p_tilde, p_tilde);
mu1[var1_idx] = mu_w;
sig1[var1_idx, var1_idx] = sigma_w;
out = append_col(mu1, sig1);
return out;
}
matrix calc_B_tilde(matrix sigma_b, vector mu_b, array[] int var2_idx, int p_tilde) {
matrix[p_tilde, p_tilde + 1] out = rep_matrix(0, p_tilde, p_tilde + 1);
vector[p_tilde] mu2 = rep_vector(0, p_tilde);
matrix[p_tilde, p_tilde] sig2 = rep_matrix(0, p_tilde, p_tilde);
mu2[var2_idx] = mu_b;
sig2[var2_idx, var2_idx] = sigma_b;
out = append_col(mu2, sig2);
return out;
}
vector twolevel_logdens(array[] vector mean_d, array[] matrix cov_d, matrix S_PW, array[] vector YX, array[] int nclus, array[] int clus_size, array[] int clus_sizes, int nclus_sizes, array[] int clus_size_ns, vector impl_Muw, matrix impl_Sigmaw, vector impl_Mub, matrix impl_Sigmab, array[] int ov_idx1, array[] int ov_idx2, array[] int within_idx, array[] int between_idx, array[] int both_idx, int p_tilde, int N_within, int N_between, int N_both){
matrix[p_tilde, p_tilde + 1] W_tilde;
matrix[p_tilde, p_tilde] W_tilde_cov;
matrix[p_tilde, p_tilde + 1] B_tilde;
matrix[p_tilde, p_tilde] B_tilde_cov;
vector[p_tilde] Mu_WB_tilde;
vector[N_between] Mu_z;
int N_wo_b = p_tilde - N_between;
vector[N_wo_b] Mu_y;
vector[N_wo_b] Mu_w;
vector[N_wo_b] Mu_b;
vector[N_between + N_wo_b] Mu_full;
matrix[N_between, N_between] Sigma_zz;
matrix[N_between, N_between] Sigma_zz_inv;
real Sigma_zz_ld;
matrix[N_wo_b, N_between] Sigma_yz;
matrix[N_wo_b, N_between] Sigma_yz_zi;
matrix[N_wo_b, N_wo_b] Sigma_b;
matrix[N_wo_b, N_wo_b] Sigma_b_z;
matrix[N_wo_b, N_wo_b] Sigma_w;
matrix[N_wo_b, N_wo_b] Sigma_w_inv;
real Sigma_w_ld;
matrix[N_wo_b, N_wo_b] Sigma_j;
matrix[N_wo_b, N_wo_b] Sigma_j_inv;
matrix[N_wo_b, N_between] Sigma_ji_yz_zi;
matrix[N_between, N_between] Vinv_11;
real Sigma_j_ld;
vector[nclus_sizes] L;
vector[nclus_sizes] B;
array[N_between] int bidx;
array[p_tilde - N_between] int notbidx;
real q_zz;
real q_yz;
real q_yyc;
vector[nclus_sizes] P;
vector[nclus_sizes] q_W;
vector[nclus_sizes] L_W;
vector[nclus_sizes] loglik;
vector[nclus_sizes] nperclus = to_vector(clus_sizes) .* to_vector(clus_size_ns);
int cluswise = 0;
int r1 = 1; // running index of rows of YX corresponding to each cluster
// 1. compute necessary vectors/matrices, like lav_mvnorm_cluster_implied22l() of lav_mvnorm_cluster.R
if (nclus[2] == nclus_sizes) cluswise = 1;
W_tilde = calc_W_tilde(impl_Sigmaw, impl_Muw, ov_idx1, p_tilde);
W_tilde_cov = block(W_tilde, 1, 2, p_tilde, p_tilde);
B_tilde = calc_B_tilde(impl_Sigmab, impl_Mub, ov_idx2, p_tilde);
B_tilde_cov = block(B_tilde, 1, 2, p_tilde, p_tilde);
Mu_WB_tilde = rep_vector(0, p_tilde);
if (N_within > 0) {
for (i in 1:N_within) {
Mu_WB_tilde[within_idx[i]] = W_tilde[within_idx[i], 1];
B_tilde[within_idx[i], 1] = 0;
}
}
if (N_both > 0) {
for (i in 1:N_both) {
Mu_WB_tilde[both_idx[i]] = B_tilde[both_idx[i], 1] + W_tilde[both_idx[i], 1];
}
}
// around line 71 of lav_mvnorm_cluster.R
if (N_between > 0) {
bidx = between_idx[1:N_between];
notbidx = between_idx[(N_between + 1):p_tilde];
Mu_z = to_vector(B_tilde[bidx, 1]);
Mu_y = Mu_WB_tilde[notbidx];
Mu_w = to_vector(W_tilde[notbidx, 1]);
Mu_b = to_vector(B_tilde[notbidx, 1]);
Sigma_zz = B_tilde_cov[bidx, bidx];
Sigma_yz = B_tilde_cov[notbidx, bidx];
Sigma_b = B_tilde_cov[notbidx, notbidx];
Sigma_w = W_tilde_cov[notbidx, notbidx];
} else {
Mu_y = Mu_WB_tilde;
Mu_w = W_tilde[,1];
Mu_b = B_tilde[,1];
Sigma_b = B_tilde_cov;
Sigma_w = W_tilde_cov;
}
// 2. compute lpdf, around line 203 of lav_mvnorm_cluster
Sigma_w_inv = inverse_spd(Sigma_w);
Sigma_w_ld = log_determinant(Sigma_w);
if (N_between > 0) {
Sigma_zz_inv = inverse_spd(Sigma_zz);
Sigma_zz_ld = log_determinant(Sigma_zz);
Sigma_yz_zi = Sigma_yz * Sigma_zz_inv;
Sigma_b_z = Sigma_b - Sigma_yz * Sigma_yz_zi';
} else {
Sigma_zz_ld = 0;
Sigma_b_z = Sigma_b;
}
Mu_full = append_row(Mu_z, Mu_y);
for (clz in 1:nclus_sizes) {
int nj = clus_sizes[clz];
matrix[N_between + N_wo_b, N_between + N_wo_b] Y2Yc = crossprod(to_matrix(mean_d[clz] - Mu_full)');
matrix[N_between, N_between] Y2Yc_zz;
matrix[N_wo_b, N_between] Y2Yc_yz;
matrix[N_wo_b, N_wo_b] Y2Yc_yy;
array[N_between] int uord_bidx;
array[N_wo_b] int uord_notbidx;
if (!cluswise) Y2Yc += cov_d[clz]; // variability between clusters of same size, will always equal 0 for clusterwise
if (N_between > 0) {
for (k in 1:N_between) {
uord_bidx[k] = k;
}
if (N_wo_b > 0) {
for (k in 1:N_wo_b) {
uord_notbidx[k] = N_between + k;
}
}
Y2Yc_zz = Y2Yc[uord_bidx, uord_bidx];
Y2Yc_yz = Y2Yc[uord_notbidx, uord_bidx];
Y2Yc_yy = Y2Yc[uord_notbidx, uord_notbidx];
} else {
Y2Yc_yy = Y2Yc;
}
Sigma_j = (nj * Sigma_b_z) + Sigma_w;
Sigma_j_inv = inverse_spd(Sigma_j); // FIXME npd exceptions for within-only
Sigma_j_ld = log_determinant(Sigma_j);
L[clz] = Sigma_zz_ld + Sigma_j_ld;
if (N_between > 0) {
Sigma_ji_yz_zi = Sigma_j_inv * Sigma_yz_zi;
Vinv_11 = Sigma_zz_inv + nj * (Sigma_yz_zi' * Sigma_ji_yz_zi);
q_zz = sum(Vinv_11 .* Y2Yc_zz);
q_yz = -nj * sum(Sigma_ji_yz_zi .* Y2Yc_yz);
} else {
q_zz = 0;
q_yz = 0;
}
q_yyc = -nj * sum(Sigma_j_inv .* Y2Yc_yy);
B[clz] = q_zz + 2 * q_yz - q_yyc;
if (cluswise) {
matrix[N_wo_b, nj] Y_j;
if (N_between > 0) {
for (i in 1:nj) {
Y_j[, i] = YX[r1 - 1 + i, notbidx] - mean_d[clz, uord_notbidx]; // would be nice to have to_matrix() here
}
} else {
for (i in 1:nj) {
Y_j[, i] = YX[r1 - 1 + i] - mean_d[clz]; // would be nice to have to_matrix() here
}
}
r1 += nj; // for next iteration through loop
q_W[clz] = sum(Sigma_w_inv .* tcrossprod(Y_j));
}
}
if (!cluswise) {
q_W = (nperclus - to_vector(clus_size_ns)) * sum(Sigma_w_inv .* S_PW);
}
L_W = (nperclus - to_vector(clus_size_ns)) * Sigma_w_ld;
loglik = -.5 * ((L .* to_vector(clus_size_ns)) + (B .* to_vector(clus_size_ns)) + q_W + L_W);
// add constant, line 300 lav_mvnorm_cluster
P = nperclus * (N_within + N_both) + to_vector(clus_size_ns) * N_between;
loglik += -.5 * (P * log(2 * pi()));
return loglik;
}
/*
Fills in the elements of a coefficient matrix containing some mix of
totally free, free subject to a sign constraint, and fixed elements
@param free_elements vector of unconstrained elements
@param skeleton matrix of the same dimensions as the output whose elements are
positive_infinity(): if output element is totally free
other: if output element is fixed to that number
@return matrix of coefficients
*/
matrix fill_matrix(vector free_elements, matrix skeleton, array[,] int eq_skeleton, int pos_start, int spos_start) {
int R = rows(skeleton);
int C = cols(skeleton);
matrix[R, C] out;
int pos = spos_start; // position of eq_skeleton
int freepos = pos_start; // position of free_elements
int eqelem = 0;
for (c in 1:C) for (r in 1:R) {
real rc = skeleton[r, c];
if (is_inf(rc)) { // free
int eq = eq_skeleton[pos, 1];
int wig = eq_skeleton[pos, 3];
if (eq == 0 || wig == 1) {
out[r,c] = free_elements[freepos];
freepos += 1;
} else {
eqelem = eq_skeleton[pos, 2];
out[r,c] = free_elements[eqelem];
}
pos += 1;
} else out[r,c] = skeleton[r, c]; // fixed, so do not bump pos
}
return out;
}
vector fill_prior(vector free_elements, array[] real pri_mean, array[,] int eq_skeleton) {
int R = dims(eq_skeleton)[1];
int eqelem = 0;
int pos = 1;
vector[num_elements(pri_mean)] out;
for (r in 1:R) {
if (pos <= num_elements(pri_mean)) {
int eq = eq_skeleton[r, 1];
int wig = eq_skeleton[r, 3];
if (eq == 0) {
out[pos] = pri_mean[pos];
pos += 1;
} else if (wig == 1) {
eqelem = eq_skeleton[r, 2];
out[pos] = free_elements[eqelem];
pos += 1;
}
}
}
return out;
}
/*
* This is a bug-free version of csr_to_dense_matrix and has the same arguments
*/
matrix to_dense_matrix(int m, int n, vector w, array[] int v, array[] int u) {
matrix[m, n] out = rep_matrix(0, m, n);
int pos = 1;
for (i in 1:m) {
int start = u[i];
int nnz = u[i + 1] - start;
for (j in 1:nnz) {
out[i, v[pos]] = w[pos];
pos += 1;
}
}
return out;
}
// sign function
int sign(real x) {
if (x > 0)
return 1;
else
return -1;
}
// sign-constrain a vector of loadings
vector sign_constrain_load(vector free_elements, int npar, array[,] int sign_mat) {
vector[npar] out;
for (i in 1:npar) {
if (sign_mat[i,1]) {
int lookupval = sign_mat[i,2];
if (free_elements[lookupval] < 0) {
out[i] = -free_elements[i];
} else {
out[i] = free_elements[i];
}
} else {
out[i] = free_elements[i];
}
}
return out;
}
// sign-constrain a vector of regressions or covariances
vector sign_constrain_reg(vector free_elements, int npar, array[,] int sign_mat, vector load_par1, vector load_par2) {
vector[npar] out;
for (i in 1:npar) {
if (sign_mat[i,1]) {
int lookupval1 = sign_mat[i,2];
int lookupval2 = sign_mat[i,3];
if (sign(load_par1[lookupval1]) * sign(load_par2[lookupval2]) < 0) {
out[i] = -free_elements[i];
} else {
out[i] = free_elements[i];
}
} else {
out[i] = free_elements[i];
}
}
return out;
}
// obtain covariance parameter vector for correlation/sd matrices
vector cor2cov(array[] matrix cormat, array[] matrix sdmat, int num_free_elements, array[] matrix matskel, array[,] int wskel, int ngrp) {
vector[num_free_elements] out;
int R = rows(to_matrix(cormat[1]));
int pos = 1; // position of eq_skeleton
int freepos = 1; // position of free_elements
for (g in 1:ngrp) {
for (c in 1:(R-1)) for (r in (c+1):R) {
if (is_inf(matskel[g,r,c])) {
if (wskel[pos,1] == 0) {
out[freepos] = sdmat[g,r,r] * sdmat[g,c,c] * cormat[g,r,c];
freepos += 1;
}
pos += 1;
}
}
}
return out;
}
// E step of EM algorithm on latent continuous space
array[] matrix estep(array[] vector YXstar, array[] vector Mu, array[] matrix Sigma, array[] int Nobs, array[,] int Obsvar, array[] int startrow, array[] int endrow, array[] int grpnum, int Np, int Ng) {
int p = dims(YXstar)[2];
array[Ng] matrix[p, p + 1] out; //mean vec + cov mat
matrix[dims(YXstar)[1], p] YXfull; // columns consistenly ordered
matrix[p, p] T2pat;
array[p] int obsidx;
int r1;
int r2;
int grpidx;
int Nmis;
for (g in 1:Ng) {
out[g] = rep_matrix(0, p, p + 1);
}
for (mm in 1:Np) {
obsidx = Obsvar[mm,];
r1 = startrow[mm];
r2 = endrow[mm];
grpidx = grpnum[mm];
Nmis = p - Nobs[mm];
if (Nobs[mm] < p) {
matrix[Nobs[mm], Nobs[mm]] Sig22 = Sigma[grpidx, obsidx[1:Nobs[mm]], obsidx[1:Nobs[mm]]];
matrix[Nmis, Nmis] Sig11 = Sigma[grpidx, obsidx[(Nobs[mm] + 1):p], obsidx[(Nobs[mm] + 1):p]];
matrix[Nmis, Nobs[mm]] Sig12 = Sigma[grpidx, obsidx[(Nobs[mm] + 1):p], obsidx[1:Nobs[mm]]];
matrix[Nobs[mm], Nobs[mm]] S22inv = inverse_spd(Sig22);
matrix[Nmis, Nmis] T2p11 = Sig11 - (Sig12 * S22inv * Sig12');
// partition into observed/missing, compute Sigmas, add to out
for (jj in r1:r2) {
vector[Nmis] ymis;
ymis = Mu[grpidx, obsidx[(Nobs[mm] + 1):p]] + (Sig12 * S22inv * (YXstar[jj, 1:Nobs[mm]] - Mu[grpidx, obsidx[1:Nobs[mm]]]));
for (kk in 1:Nobs[mm]) {
YXfull[jj, obsidx[kk]] = YXstar[jj, kk];
}
for (kk in (Nobs[mm] + 1):p) {
YXfull[jj, obsidx[kk]] = ymis[kk - Nobs[mm]];
}
}
T2pat = crossprod(YXfull[r1:r2,]);
// correction for missing cells/conditional covariances
for (jj in 1:Nmis) {
for (kk in jj:Nmis) {
T2pat[obsidx[Nobs[mm] + jj], obsidx[Nobs[mm] + kk]] = T2pat[obsidx[Nobs[mm] + jj], obsidx[Nobs[mm] + kk]] + (r2 - r1 + 1) * T2p11[jj, kk];
if (kk > jj) {
T2pat[obsidx[Nobs[mm] + kk], obsidx[Nobs[mm] + jj]] = T2pat[obsidx[Nobs[mm] + jj], obsidx[Nobs[mm] + kk]];
}
}
}
} else {
// complete data
for (jj in r1:r2) {
for (kk in 1:Nobs[mm]) {
YXfull[jj, obsidx[kk]] = YXstar[jj, kk];
}
}
T2pat = crossprod(YXfull[r1:r2,]);
}
for (i in 1:p) {
out[grpidx,i,1] += sum(YXfull[r1:r2,i]);
}
out[grpidx,,2:(p+1)] += T2pat;
}
return out;
}
matrix sig_inv_update(matrix Sigmainv, array[] int obsidx, int Nobs, int np, real logdet) {
matrix[Nobs + 1, Nobs + 1] out = rep_matrix(0, Nobs + 1, Nobs + 1);
int nrm = np - Nobs;
matrix[nrm, nrm] H;
matrix[nrm, Nobs] A;
if (nrm == 0) {
out[1:Nobs, 1:Nobs] = Sigmainv;
out[Nobs + 1, Nobs + 1] = logdet;
} else {
H = Sigmainv[obsidx[(Nobs + 1):np], obsidx[(Nobs + 1):np]];
A = Sigmainv[obsidx[(Nobs + 1):np], obsidx[1:Nobs]];
out[1:Nobs, 1:Nobs] = Sigmainv[obsidx[1:Nobs], obsidx[1:Nobs]] - A' * mdivide_left_spd(H, A);
out[Nobs + 1, Nobs + 1] = logdet + log_determinant(H);
}
return out;
}
real multi_normal_suff(vector xbar, matrix S, vector Mu, matrix Supdate, int N) {
int Nobs = dims(S)[1];
real out;
// using elementwise multiplication + sum here for efficiency
out = -.5 * N * ( sum(Supdate[1:Nobs, 1:Nobs] .* (S + (xbar - Mu) * (xbar - Mu)')) + Supdate[Nobs + 1, Nobs + 1] + Nobs * log(2 * pi()) );
if(is_nan(out) || out == positive_infinity()) out = negative_infinity();
return out;
}
// compute mean vectors and cov matrices for a single group (two-level models)
array[] vector calc_mean_vecs(array[] vector YXstar, array[] vector mean_d, array[] int nclus, array[] int Xvar, array[] int Xbetvar, int Nx, int Nx_between, int p_tilde) {
vector[Nx] ov_mean = rep_vector(0, Nx);
vector[Nx_between] ov_mean_d = rep_vector(0, Nx_between);
int nr = dims(YXstar)[1];
array[2] vector[p_tilde] out;
for (i in 1:2) out[i] = rep_vector(0, p_tilde);
if (Nx > 0) {
for (i in 1:nr) {
ov_mean += YXstar[i, Xvar[1:Nx]];
}
ov_mean *= pow(nclus[1], -1);
out[1, 1:Nx] = ov_mean;
}
if (Nx_between > 0) {
for (cc in 1:nclus[2]) {
ov_mean_d += mean_d[cc, 1:Nx_between];
}
ov_mean_d *= pow(nclus[2], -1);
out[2, 1:Nx_between] = ov_mean_d;
}
return out;
}
array[] matrix calc_cov_mats(array[] vector YXstar, array[] vector mean_d, array[] vector mean_vecs, array[] int nclus, array[] int Xvar, array[] int Xbetvar, int Nx, int Nx_between, int p_tilde) {
matrix[Nx_between, Nx_between] cov_mean_d = rep_matrix(0, Nx_between, Nx_between);
matrix[Nx, Nx] cov_w = rep_matrix(0, Nx, Nx);
matrix[Nx, Nx] cov_w_inv;
int nr = dims(YXstar)[1];
array[3] matrix[p_tilde, p_tilde] out;
for (i in 1:3) out[i] = rep_matrix(0, p_tilde, p_tilde);
if (Nx > 0) {
for (i in 1:nr) {
cov_w += tcrossprod(to_matrix(YXstar[i, Xvar[1:Nx]] - mean_vecs[1, 1:Nx]));
}
cov_w *= pow(nclus[1], -1);
cov_w_inv[1:Nx, 1:Nx] = inverse_spd(cov_w);
out[2, 1:Nx, 1:Nx] = cov_w;
out[3, 1:Nx, 1:Nx] = cov_w_inv;
out[3, Nx + 1, Nx + 1] = log_determinant(cov_w); // need log_determinant for multi_normal_suff
}
if (Nx_between > 0) {
for (cc in 1:nclus[2]) {
cov_mean_d += tcrossprod(to_matrix(mean_d[cc, 1:Nx_between] - mean_vecs[2, 1:Nx_between]));
}
cov_mean_d *= pow(nclus[2], -1);
out[1, 1:Nx_between, 1:Nx_between] = cov_mean_d;
}
return out;
}
// compute log_lik of fixed.x variables for a single group (two-level models)
vector calc_log_lik_x(array[] vector mean_d, vector ov_mean_d, matrix cov_mean_d, matrix cov_w, matrix cov_w_inv, array[] int nclus, array[] int cluster_size, array[] int Xvar, array[] int Xbetvar, int Nx, int Nx_between) {
vector[nclus[2]] out = rep_vector(0, nclus[2]);
for (cc in 1:nclus[2]) {
if (Nx > 0) {
out[cc] += multi_normal_suff(mean_d[cc, Xvar[1:Nx]], cov_w[1:Nx, 1:Nx], mean_d[cc, Xvar[1:Nx]], cov_w_inv[1:(Nx + 1), 1:(Nx + 1)], cluster_size[cc]);
}
if (Nx_between > 0) {
out[cc] += multi_normal_lpdf(mean_d[cc, 1:Nx_between] | ov_mean_d[1:Nx_between], cov_mean_d[1:Nx_between, 1:Nx_between]);
}
}
return out;
}
// fill covariance matrix with blocks
array[] matrix fill_cov(array[] matrix covmat, array[,] int blkse, array[] int nblk,
array[] matrix mat_1, array[] matrix mat_2, array[] matrix mat_3,
array[] matrix mat_4, array[] matrix mat_5) {
array[dims(covmat)[1]] matrix[dims(covmat)[2], dims(covmat)[3]] out = covmat;
for (k in 1:sum(nblk)) {
int blkidx = blkse[k, 6];
int arrayidx = blkse[k, 5];
int blkgrp = blkse[k, 4];
int srow = blkse[k, 1];
int erow = blkse[k, 2];
if (arrayidx == 1) {
out[blkgrp, srow:erow, srow:erow] = mat_1[blkidx];
} else if (arrayidx == 2) {
out[blkgrp, srow:erow, srow:erow] = mat_2[blkidx];
} else if (arrayidx == 3) {
out[blkgrp, srow:erow, srow:erow] = mat_3[blkidx];
} else if (arrayidx == 4) {
out[blkgrp, srow:erow, srow:erow] = mat_4[blkidx];
} else {
out[blkgrp, srow:erow, srow:erow] = mat_5[blkidx];
}
}
return out;
}
}
data {
// see https://books.google.com/books?id=9AC-s50RjacC&lpg=PP1&dq=LISREL&pg=PA2#v=onepage&q=LISREL&f=false
int<lower=0> p; // number of manifest response variables
int<lower=0> p_c; // number of manifest level 2 variables
int<lower=0> q; // number of manifest predictors
int<lower=0> m; // number of latent endogenous variables
int<lower=0> m_c; // number of latent level 2 variables
int<lower=0> n; // number of latent exogenous variables
int<lower=1> Ng; // number of groups
int<lower=0, upper=1> missing; // are there missing values?
int<lower=0, upper=1> save_lvs; // should we save lvs?
int<lower=1> Np; // number of group-by-missing patterns combos
array[Ng] int<lower=1> N; // number of observations per group
array[Np] int<lower=1> Nobs; // number of observed variables in each missing pattern
array[Np] int<lower=0> Nordobs; // number of ordinal observed variables in each missing pattern
array[Np, p + q] int<lower=0> Obsvar; // indexing of observed variables
int<lower=1> Ntot; // number of observations across all groups
array[Np] int<lower=1> startrow; // starting row for each missing pattern
array[Np] int<lower=1,upper=Ntot> endrow; // ending row for each missing pattern
array[Np] int<lower=1,upper=Ng> grpnum; // group number for each row of data
int<lower=0,upper=1> wigind; // do any parameters have approx equality constraint ('wiggle')?
int<lower=0, upper=1> has_data; // are the raw data on y and x available?
int<lower=0, upper=1> ord; // are there any ordinal variables?
int<lower=0, upper=1> multilev; // is this a multilevel dataset?
int<lower=0> Nord; // how many ordinal variables?
array[Nord] int<lower=0> ordidx; // indexing of ordinal variables
array[Np, Nord] int<lower=0> OrdObsvar; // indexing of observed ordinal variables in YXo
int<lower=0> Noent; // how many observed entries of ordinal variables (for data augmentation)
array[p + q - Nord] int<lower=0> contidx; // indexing of continuous variables
array[Nord] int<lower=1> nlevs; // how many levels does each ordinal variable have
array[Ng, 2] int<lower=1> nclus; // number of level 1 + level 2 observations
int<lower=0> p_tilde; // total number of variables
array[Ntot] vector[multilev ? p_tilde : p + q - Nord] YX; // continuous data
array[Ntot, Nord] int YXo; // ordinal data
array[Np] int<lower=0> Nx; // number of fixed.x variables (within)
array[Np] int<lower=0> Nx_between; // number of fixed.x variables (between)
int<lower=0, upper=1> use_cov;
int<lower=0, upper=1> pri_only;
int<lower=0> emiter; // number of em iterations for saturated model in ppp (missing data only)
int<lower=0, upper=1> use_suff; // should we compute likelihood via mvn sufficient stats?
int<lower=0, upper=1> do_test; // should we do everything in generated quantities?
array[Np] vector[multilev ? p_tilde : p + q - Nord] YXbar; // sample means of continuous manifest variables
array[Np] matrix[multilev ? (p_tilde + 1) : (p + q - Nord + 1), multilev ? (p_tilde + 1) : (p + q - Nord + 1)] S; // sample covariance matrix among all continuous manifest variables NB!! multiply by (N-1) to use wishart lpdf!!
array[sum(nclus[,2])] int<lower=1> cluster_size; // number of obs per cluster
array[Ng] int<lower=1> ncluster_sizes; // number of unique cluster sizes
array[sum(ncluster_sizes)] int<lower=1> cluster_sizes; // unique cluster sizes
array[sum(ncluster_sizes)] int<lower=1> cluster_size_ns; // number of clusters of each size
array[Np, multilev ? p_tilde : p + q] int<lower=0> Xvar; // indexing of fixed.x variables (within)
array[Np, multilev ? p_tilde : p + q] int<lower=0> Xdatvar; // indexing of fixed.x in data (differs from Xvar when missing)
array[Np, multilev ? p_tilde : p + q] int<lower=0> Xbetvar; // indexing of fixed.x variables (between)
array[sum(ncluster_sizes)] vector[p_tilde] mean_d; // sample means by unique cluster size
array[sum(ncluster_sizes)] matrix[p_tilde, p_tilde] cov_d; // sample covariances by unique cluster size
array[Ng] matrix[p_tilde, p_tilde] cov_w; // observed "within" covariance matrix
array[sum(nclus[,2])] vector[p_tilde] mean_d_full; // sample means/covs by cluster, for clusterwise log-densities
array[sum(nclus[,2])] matrix[p_tilde, p_tilde] cov_d_full;
array[Ng] vector[p_tilde] xbar_w; // data estimates of within/between means/covs (for saturated logl)
array[Ng] vector[p_tilde] xbar_b;
array[Ng] matrix[p_tilde, p_tilde] cov_b;
array[Ng] real gs; // group size constant, for computation of saturated logl
int N_within; // number of within variables
int N_between; // number of between variables
int N_both; // number of variables at both levels
array[2] int N_lev; // number of observed variables at each level
array[N_within] int within_idx;
array[p_tilde] int between_idx; // between indexing, followed by within/both
array[N_lev[1]] int ov_idx1;
array[N_lev[2]] int ov_idx2;
array[N_both] int both_idx;
vector[multilev ? sum(ncluster_sizes) : Ng] log_lik_x; // ll of fixed x variables by unique cluster size
vector[multilev ? sum(nclus[,2]) : Ng] log_lik_x_full; // ll of fixed x variables by cluster
/* sparse matrix representations of skeletons of coefficient matrices,
which is not that interesting but necessary because you cannot pass
missing values into the data block of a Stan program from R */
int<lower=0> len_w1; // max number of free elements in Lambda_y per grp
array[Ng] int<lower=0> wg1; // number of free elements in Lambda_y per grp
array[Ng] vector[len_w1] w1; // values of free elements in Lambda_y
array[Ng, len_w1] int<lower=1> v1; // index of free elements in Lambda_y
array[Ng, p + 1] int<lower=1> u1; // index of free elements in Lambda_y
array[sum(wg1), 3] int<lower=0> w1skel;
array[sum(wg1), 2] int<lower=0> lam_y_sign;
int<lower=0> len_lam_y; // number of free elements minus equality constraints
array[len_lam_y] real lambda_y_mn; // prior
array[len_lam_y] real<lower=0> lambda_y_sd;
// same things but for B
int<lower=0> len_w4;
array[Ng] int<lower=0> wg4;
array[Ng] vector[len_w4] w4;
array[Ng, len_w4] int<lower=1> v4;
array[Ng, m + 1] int<lower=1> u4;
array[sum(wg4), 3] int<lower=0> w4skel;
array[sum(wg4), 3] int<lower=0> b_sign;
int<lower=0> len_b;
array[len_b] real b_mn;
array[len_b] real<lower=0> b_sd;
// same things but for diag(Theta)
int<lower=0> len_w5;
array[Ng] int<lower=0> wg5;
array[Ng] vector[len_w5] w5;
array[Ng, len_w5] int<lower=1> v5;
array[Ng, p + 1] int<lower=1> u5;
array[sum(wg5), 3] int<lower=0> w5skel;
int<lower=0> len_thet_sd;
array[len_thet_sd] real<lower=0> theta_sd_shape;
array[len_thet_sd] real<lower=0> theta_sd_rate;
int<lower=-2, upper=2> theta_pow;
// same things but for Theta_r
int<lower=0> len_w7;
array[Ng] int<lower=0> wg7;
array[Ng] vector[len_w7] w7;
array[Ng, len_w7] int<lower=1> v7;
array[Ng, p + 1] int<lower=1> u7;
array[sum(wg7), 3] int<lower=0> w7skel;
int<lower=0> len_thet_r;
array[len_thet_r] real<lower=0> theta_r_alpha;
array[len_thet_r] real<lower=0> theta_r_beta;
// same things but for Psi
int<lower=0> len_w9;
array[Ng] int<lower=0> wg9;
array[Ng] vector[len_w9] w9;
array[Ng, len_w9] int<lower=1> v9;
array[Ng, m + 1] int<lower=1> u9;
array[sum(wg9), 3] int<lower=0> w9skel;
int<lower=0> len_psi_sd;
array[len_psi_sd] real<lower=0> psi_sd_shape;
array[len_psi_sd] real<lower=0> psi_sd_rate;
int<lower=-2,upper=2> psi_pow;
// same things but for Psi_r
int<lower=0> len_w10;
array[Ng] int<lower=0> wg10;
array[Ng] vector[len_w10] w10;
array[Ng, len_w10] int<lower=1> v10;
array[Ng, m + 1] int<lower=1> u10;
array[sum(wg10), 3] int<lower=0> w10skel;
array[sum(wg10), 3] int<lower=0> psi_r_sign;
int<lower=0> len_psi_r;
array[len_psi_r] real<lower=0> psi_r_alpha;
array[len_psi_r] real<lower=0> psi_r_beta;
// for blocks within Psi_r that receive lkj
array[5] int<lower=0> nblk;
array[5] int<lower=3> psidims;
array[sum(nblk), 7] int<lower=0> blkse;
int<lower=0> len_w11;
array[Ng] int<lower=0> wg11;
array[Ng] vector[len_w11] w11;
array[Ng, len_w11] int<lower=1> v11;
array[Ng, m + 1] int<lower=1> u11;
array[sum(wg11), 3] int<lower=0> w11skel;
// same things but for Nu
int<lower=0> len_w13;
array[Ng] int<lower=0> wg13;
array[Ng] vector[len_w13] w13;
array[Ng, len_w13] int<lower=1> v13;
array[Ng, use_cov ? 1 : p + q + 1] int<lower=1> u13;
array[sum(wg13), 3] int<lower=0> w13skel;
int<lower=0> len_nu;
array[len_nu] real nu_mn;
array[len_nu] real<lower=0> nu_sd;
// same things but for Alpha
int<lower=0> len_w14;
array[Ng] int<lower=0> wg14;
array[Ng] vector[len_w14] w14;
array[Ng, len_w14] int<lower=0> v14;
array[Ng, use_cov ? 1 : m + n + 1] int<lower=1> u14;
array[sum(wg14), 3] int<lower=0> w14skel;
int<lower=0> len_alph;
array[len_alph] real alpha_mn;
array[len_alph] real<lower=0> alpha_sd;
// same things but for Tau
int<lower=0> len_w15;
array[Ng] int<lower=0> wg15;
array[Ng] vector[len_w15] w15;
array[Ng, len_w15] int<lower=0> v15;
array[Ng, sum(nlevs) - Nord + 1] int<lower=1> u15;
array[sum(wg15), 3] int<lower=0> w15skel;
int<lower=0> len_tau;
array[len_tau] real tau_mn;
array[len_tau] real<lower=0> tau_sd;
// Level 2 matrices start here!!
// Lambda
int<lower=0> len_w1_c;
array[Ng] int<lower=0> wg1_c;
array[Ng] vector[len_w1_c] w1_c;
array[Ng, len_w1_c] int<lower=1> v1_c;
array[Ng, p_c + 1] int<lower=1> u1_c;
array[sum(wg1_c), 3] int<lower=0> w1skel_c;
array[sum(wg1_c), 2] int<lower=0> lam_y_sign_c;
int<lower=0> len_lam_y_c;
array[len_lam_y_c] real lambda_y_mn_c;
array[len_lam_y_c] real<lower=0> lambda_y_sd_c;
// same things but for B
int<lower=0> len_w4_c;
array[Ng] int<lower=0> wg4_c;
array[Ng] vector[len_w4_c] w4_c;
array[Ng, len_w4_c] int<lower=1> v4_c;
array[Ng, m_c + 1] int<lower=1> u4_c;
array[sum(wg4_c), 3] int<lower=0> w4skel_c;
array[sum(wg4_c), 3] int<lower=0> b_sign_c;
int<lower=0> len_b_c;
array[len_b_c] real b_mn_c;
array[len_b_c] real<lower=0> b_sd_c;
// same things but for diag(Theta)
int<lower=0> len_w5_c;
array[Ng] int<lower=0> wg5_c;
array[Ng] vector[len_w5_c] w5_c;
array[Ng, len_w5_c] int<lower=1> v5_c;
array[Ng, p_c + 1] int<lower=1> u5_c;
array[sum(wg5_c), 3] int<lower=0> w5skel_c;
int<lower=0> len_thet_sd_c;
array[len_thet_sd_c] real<lower=0> theta_sd_shape_c;
array[len_thet_sd_c] real<lower=0> theta_sd_rate_c;
int<lower=-2, upper=2> theta_pow_c;
// same things but for Theta_r
int<lower=0> len_w7_c;
array[Ng] int<lower=0> wg7_c;
array[Ng] vector[len_w7_c] w7_c;
array[Ng, len_w7_c] int<lower=1> v7_c;
array[Ng, p_c + 1] int<lower=1> u7_c;
array[sum(wg7_c), 3] int<lower=0> w7skel_c;
int<lower=0> len_thet_r_c;
array[len_thet_r_c] real<lower=0> theta_r_alpha_c;
array[len_thet_r_c] real<lower=0> theta_r_beta_c;
// same things but for Psi
int<lower=0> len_w9_c;
array[Ng] int<lower=0> wg9_c;
array[Ng] vector[len_w9_c] w9_c;
array[Ng, len_w9_c] int<lower=1> v9_c;
array[Ng, m_c + 1] int<lower=1> u9_c;
array[sum(wg9_c), 3] int<lower=0> w9skel_c;
int<lower=0> len_psi_sd_c;
array[len_psi_sd_c] real<lower=0> psi_sd_shape_c;
array[len_psi_sd_c] real<lower=0> psi_sd_rate_c;
int<lower=-2,upper=2> psi_pow_c;
// same things but for Psi_r
int<lower=0> len_w10_c;
array[Ng] int<lower=0> wg10_c;
array[Ng] vector[len_w10_c] w10_c;
array[Ng, len_w10_c] int<lower=1> v10_c;
array[Ng, m_c + 1] int<lower=1> u10_c;
array[sum(wg10_c), 3] int<lower=0> w10skel_c;
array[sum(wg10_c), 3] int<lower=0> psi_r_sign_c;
int<lower=0> len_psi_r_c;
array[len_psi_r_c] real<lower=0> psi_r_alpha_c;
array[len_psi_r_c] real<lower=0> psi_r_beta_c;
// for blocks within Psi_r that receive lkj
array[5] int<lower=0> nblk_c;
array[5] int<lower=3> psidims_c;
array[sum(nblk_c), 7] int<lower=0> blkse_c;
int<lower=0> len_w11_c;
array[Ng] int<lower=0> wg11_c;
array[Ng] vector[len_w11_c] w11_c;
array[Ng, len_w11_c] int<lower=1> v11_c;
array[Ng, m_c + 1] int<lower=1> u11_c;
array[sum(wg11_c), 3] int<lower=0> w11skel_c;
// same things but for Nu
int<lower=0> len_w13_c;
array[Ng] int<lower=0> wg13_c;
array[Ng] vector[len_w13_c] w13_c;
array[Ng, len_w13_c] int<lower=1> v13_c;
array[Ng, p_c + 1] int<lower=1> u13_c;
array[sum(wg13_c), 3] int<lower=0> w13skel_c;
int<lower=0> len_nu_c;
array[len_nu_c] real nu_mn_c;
array[len_nu_c] real<lower=0> nu_sd_c;
// same things but for Alpha
int<lower=0> len_w14_c;
array[Ng] int<lower=0> wg14_c;
array[Ng] vector[len_w14_c] w14_c;
array[Ng, len_w14_c] int<lower=0> v14_c;
array[Ng, m_c + 1] int<lower=1> u14_c;
array[sum(wg14_c), 3] int<lower=0> w14skel_c;
int<lower=0> len_alph_c;
array[len_alph_c] real alpha_mn_c;
array[len_alph_c] real<lower=0> alpha_sd_c;
}
transformed data { // (re)construct skeleton matrices in Stan (not that interesting)
array[Ng] matrix[p, m] Lambda_y_skeleton;
array[Ng] matrix[m, m] B_skeleton;
array[Ng] matrix[p, p] Theta_skeleton;
array[Ng] matrix[p, p] Theta_r_skeleton;
array[Ng] matrix[m, m] Psi_skeleton;
array[Ng] matrix[m, m] Psi_r_skeleton;
array[Ng] matrix[m, m] Psi_r_skeleton_f;
array[Ng] matrix[p, 1] Nu_skeleton;
array[Ng] matrix[m, 1] Alpha_skeleton;
array[Ng] matrix[sum(nlevs) - Nord, 1] Tau_skeleton;
array[Np] vector[ord ? 0 : (p + q)] YXbarstar;
array[Np] matrix[ord ? 0 : (p + q), ord ? 0 : (p + q)] Sstar;
array[Ng] matrix[p_c, m_c] Lambda_y_skeleton_c;
array[Ng] matrix[m_c, m_c] B_skeleton_c;
array[Ng] matrix[p_c, p_c] Theta_skeleton_c;
array[Ng] matrix[p_c, p_c] Theta_r_skeleton_c;
array[Ng] matrix[m_c, m_c] Psi_skeleton_c;
array[Ng] matrix[m_c, m_c] Psi_r_skeleton_c;
array[Ng] matrix[m_c, m_c] Psi_r_skeleton_f_c;
array[Ng] matrix[p_c, 1] Nu_skeleton_c;
array[Ng] matrix[m_c, 1] Alpha_skeleton_c;
matrix[m, m] I = diag_matrix(rep_vector(1, m));
matrix[m_c, m_c] I_c = diag_matrix(rep_vector(1, m_c));
int Ncont = p + q - Nord;
array[max(nclus[,2]) > 1 ? max(nclus[,2]) : 0] int<lower = 0> intone;
array[Ng,2] int g_start1;
array[Ng,2] int g_start4;
array[Ng,2] int g_start5;
array[Ng,2] int g_start7;
array[Ng,2] int g_start9;
array[Ng,2] int g_start10;
array[Ng,2] int g_start13;
array[Ng,2] int g_start14;
array[Ng,2] int g_start15;
array[Ng,2] int g_start1_c;
array[Ng,2] int g_start4_c;
array[Ng,2] int g_start5_c;
array[Ng,2] int g_start7_c;
array[Ng,2] int g_start9_c;
array[Ng,2] int g_start10_c;
array[Ng,2] int g_start13_c;
array[Ng,2] int g_start14_c;
array[15] int len_free;
array[15] int pos;
array[15] int len_free_c;
array[15] int pos_c;
for (i in 1:15) {
len_free[i] = 0;
pos[i] = 1;
len_free_c[i] = 0;
pos_c[i] = 1;
}
for (g in 1:Ng) {
Lambda_y_skeleton[g] = to_dense_matrix(p, m, w1[g], v1[g,], u1[g,]);
B_skeleton[g] = to_dense_matrix(m, m, w4[g], v4[g,], u4[g,]);
Theta_skeleton[g] = to_dense_matrix(p, p, w5[g], v5[g,], u5[g,]);
Theta_r_skeleton[g] = to_dense_matrix(p, p, w7[g], v7[g,], u7[g,]);
Psi_skeleton[g] = to_dense_matrix(m, m, w9[g], v9[g,], u9[g,]);
Psi_r_skeleton[g] = to_dense_matrix(m, m, w10[g], v10[g,], u10[g,]);
Psi_r_skeleton_f[g] = to_dense_matrix(m, m, w11[g], v11[g,], u11[g,]);
if (!use_cov) {
Nu_skeleton[g] = to_dense_matrix((p + q), 1, w13[g], v13[g,], u13[g,]);
Alpha_skeleton[g] = to_dense_matrix((m + n), 1, w14[g], v14[g,], u14[g,]);
}
Tau_skeleton[g] = to_dense_matrix(sum(nlevs) - Nord, 1, w15[g], v15[g,], u15[g,]);
Lambda_y_skeleton_c[g] = to_dense_matrix(p_c, m_c, w1_c[g], v1_c[g,], u1_c[g,]);
B_skeleton_c[g] = to_dense_matrix(m_c, m_c, w4_c[g], v4_c[g,], u4_c[g,]);
Theta_skeleton_c[g] = to_dense_matrix(p_c, p_c, w5_c[g], v5_c[g,], u5_c[g,]);
Theta_r_skeleton_c[g] = to_dense_matrix(p_c, p_c, w7_c[g], v7_c[g,], u7_c[g,]);
Psi_skeleton_c[g] = to_dense_matrix(m_c, m_c, w9_c[g], v9_c[g,], u9_c[g,]);
Psi_r_skeleton_c[g] = to_dense_matrix(m_c, m_c, w10_c[g], v10_c[g,], u10_c[g,]);
Psi_r_skeleton_f_c[g] = to_dense_matrix(m_c, m_c, w11_c[g], v11_c[g,], u11_c[g,]);
Nu_skeleton_c[g] = to_dense_matrix(p_c, 1, w13_c[g], v13_c[g,], u13_c[g,]);
Alpha_skeleton_c[g] = to_dense_matrix(m_c, 1, w14_c[g], v14_c[g,], u14_c[g,]);
// count free elements in Lambda_y_skeleton
g_start1[g,1] = len_free[1] + 1;
g_start1[g,2] = pos[1];
for (i in 1:p) {
for (j in 1:m) {
if (is_inf(Lambda_y_skeleton[g,i,j])) {
if (w1skel[pos[1],2] == 0 || w1skel[pos[1],3] == 1) len_free[1] += 1;
pos[1] += 1;
}
}
}
// same thing but for B_skeleton
g_start4[g,1] = len_free[4] + 1;
g_start4[g,2] = pos[4];
for (i in 1:m) {
for (j in 1:m) {
if (is_inf(B_skeleton[g,i,j])) {
if (w4skel[pos[4],2] == 0 || w4skel[pos[4],3] == 1) len_free[4] += 1;
pos[4] += 1;
}
}
}
// same thing but for Theta_skeleton
g_start5[g,1] = len_free[5] + 1;
g_start5[g,2] = pos[5];
for (i in 1:p) {
if (is_inf(Theta_skeleton[g,i,i])) {
if (w5skel[pos[5],2] == 0 || w5skel[pos[5],3] == 1) len_free[5] += 1;
pos[5] += 1;
}
}
// same thing but for Theta_r_skeleton
g_start7[g,1] = len_free[7] + 1;
g_start7[g,2] = pos[7];
for (i in 1:(p-1)) {
for (j in (i+1):p) {
if (is_inf(Theta_r_skeleton[g,j,i])) {
if (w7skel[pos[7],2] == 0 || w7skel[pos[7],3] == 1) len_free[7] += 1;
pos[7] += 1;
}
}
}
// same thing but for Psi_skeleton
g_start9[g,1] = len_free[9] + 1;
g_start9[g,2] = pos[9];
for (i in 1:m) {
if (is_inf(Psi_skeleton[g,i,i])) {
if (w9skel[pos[9],2] == 0 || w9skel[pos[9],3] == 1) len_free[9] += 1;
pos[9] += 1;
}
}
// same thing but for Psi_r_skeleton
g_start10[g,1] = len_free[10] + 1;
g_start10[g,2] = pos[10];
for (i in 1:(m-1)) {
for (j in (i+1):m) {
if (is_inf(Psi_r_skeleton[g,j,i])) {
if (w10skel[pos[10],2] == 0 || w10skel[pos[10],3] == 1) len_free[10] += 1;
pos[10] += 1;
}
if (is_inf(Psi_r_skeleton_f[g,j,i])) {
if (w11skel[pos[11],2] == 0 || w11skel[pos[11],3] == 1) len_free[11] += 1;
pos[11] += 1;
}
}
}
if (!use_cov) {
// same thing but for Nu_skeleton
// pos = len_free13 + 1;
g_start13[g,1] = len_free[13] + 1;
g_start13[g,2] = pos[13];
for (i in 1:(p+q)) {
if (is_inf(Nu_skeleton[g,i,1])) {
if (w13skel[pos[13],2] == 0 || w13skel[pos[13],3] == 1) len_free[13] += 1;
pos[13] += 1;
}
}
// same thing but for Alpha_skeleton
g_start14[g,1] = len_free[14] + 1;
g_start14[g,2] = pos[14];
for (i in 1:(m+n)) {
if (is_inf(Alpha_skeleton[g,i,1])) {
if (w14skel[pos[14],2] == 0 || w14skel[pos[14],3] == 1) len_free[14] += 1;
pos[14] += 1;
}
}
}
// same thing but for Tau_skeleton
g_start15[g,1] = len_free[15] + 1;
g_start15[g,2] = pos[15];
for (i in 1:(sum(nlevs) - Nord)) {
if (is_inf(Tau_skeleton[g,i,1])) {
if (w15skel[pos[15],2] == 0 || w15skel[pos[15],3] == 1) len_free[15] += 1;
pos[15] += 1;
}
}
// now level 2
// count free elements in Lambda_y_skeleton
g_start1_c[g,1] = len_free_c[1] + 1;
g_start1_c[g,2] = pos_c[1];
for (i in 1:p_c) {
for (j in 1:m_c) {
if (is_inf(Lambda_y_skeleton_c[g,i,j])) {
if (w1skel_c[pos_c[1],2] == 0 || w1skel_c[pos_c[1],3] == 1) len_free_c[1] += 1;
pos_c[1] += 1;
}
}
}
// same thing but for B_skeleton
g_start4_c[g,1] = len_free_c[4] + 1;
g_start4_c[g,2] = pos_c[4];
for (i in 1:m_c) {
for (j in 1:m_c) {
if (is_inf(B_skeleton_c[g,i,j])) {
if (w4skel_c[pos_c[4],2] == 0 || w4skel_c[pos_c[4],3] == 1) len_free_c[4] += 1;
pos_c[4] += 1;
}
}
}
// same thing but for Theta_skeleton
g_start5_c[g,1] = len_free_c[5] + 1;
g_start5_c[g,2] = pos_c[5];
for (i in 1:p_c) {
if (is_inf(Theta_skeleton_c[g,i,i])) {
if (w5skel_c[pos_c[5],2] == 0 || w5skel_c[pos_c[5],3] == 1) len_free_c[5] += 1;
pos_c[5] += 1;
}
}
// same thing but for Theta_r_skeleton
g_start7_c[g,1] = len_free_c[7] + 1;
g_start7_c[g,2] = pos_c[7];
for (i in 1:(p_c-1)) {
for (j in (i+1):p_c) {
if (is_inf(Theta_r_skeleton_c[g,j,i])) {
if (w7skel_c[pos_c[7],2] == 0 || w7skel_c[pos_c[7],3] == 1) len_free_c[7] += 1;
pos_c[7] += 1;
}
}
}
// same thing but for Psi_skeleton
g_start9_c[g,1] = len_free_c[9] + 1;
g_start9_c[g,2] = pos_c[9];
for (i in 1:m_c) {
if (is_inf(Psi_skeleton_c[g,i,i])) {
if (w9skel_c[pos_c[9],2] == 0 || w9skel_c[pos_c[9],3] == 1) len_free_c[9] += 1;
pos_c[9] += 1;
}
}
// same thing but for Psi_r_skeleton
g_start10_c[g,1] = len_free_c[10] + 1;
g_start10_c[g,2] = pos_c[10];
for (i in 1:(m_c-1)) {
for (j in (i+1):m_c) {
if (is_inf(Psi_r_skeleton_c[g,j,i])) {
if (w10skel_c[pos_c[10],2] == 0 || w10skel_c[pos_c[10],3] == 1) len_free_c[10] += 1;
pos_c[10] += 1;
}
if (is_inf(Psi_r_skeleton_f_c[g,j,i])) {
if (w11skel_c[pos_c[11],2] == 0 || w11skel_c[pos_c[11],3] == 1) len_free_c[11] += 1;
pos_c[11] += 1;
}
}
}
// same thing but for Nu_skeleton
// pos = len_free13 + 1;
g_start13_c[g,1] = len_free_c[13] + 1;
g_start13_c[g,2] = pos_c[13];
for (i in 1:p_c) {
if (is_inf(Nu_skeleton_c[g,i,1])) {
if (w13skel_c[pos_c[13],2] == 0 || w13skel_c[pos_c[13],3] == 1) len_free_c[13] += 1;
pos_c[13] += 1;
}
}
// same thing but for Alpha_skeleton
g_start14_c[g,1] = len_free_c[14] + 1;
g_start14_c[g,2] = pos_c[14];
for (i in 1:m_c) {
if (is_inf(Alpha_skeleton_c[g,i,1])) {
if (w14skel_c[pos_c[14],2] == 0 || w14skel_c[pos_c[14],3] == 1) len_free_c[14] += 1;
pos_c[14] += 1;
}
}
}
// for clusterwise loglik computations
if (max(nclus[,2]) > 1) for (i in 1:max(nclus[,2])) intone[i] = 1;
if (!ord && (use_suff || use_cov)) {
// sufficient stat matrices by pattern, moved to left for missing
for (patt in 1:Np) {
Sstar[patt] = rep_matrix(0, p + q, p + q);
Sstar[patt, 1:Nobs[patt], 1:Nobs[patt]] = S[patt, Obsvar[patt, 1:Nobs[patt]], Obsvar[patt, 1:Nobs[patt]]];
for (j in 1:Nobs[patt]) {
YXbarstar[patt,j] = YXbar[patt, Obsvar[patt,j]];
}
}
}
}
parameters {
// free elements (possibly with inequality constraints) for coefficient matrices
vector[len_free[1]] Lambda_y_free;
vector[len_free[4]] B_free;
vector<lower=0>[len_free[5]] Theta_sd_free;
vector<lower=-1,upper=1>[len_free[7]] Theta_r_free; // to use beta prior
vector<lower=0>[len_free[9]] Psi_sd_free;
array[nblk[1]] corr_matrix[psidims[1]] Psi_r_mat_1;
array[nblk[2]] corr_matrix[psidims[2]] Psi_r_mat_2;
array[nblk[3]] corr_matrix[psidims[3]] Psi_r_mat_3;
array[nblk[4]] corr_matrix[psidims[4]] Psi_r_mat_4;
array[nblk[5]] corr_matrix[psidims[5]] Psi_r_mat_5;
vector<lower=-1,upper=1>[len_free[10]] Psi_r_free;
vector[len_free[13]] Nu_free;
vector[len_free[14]] Alpha_free;
vector[len_free[15]] Tau_ufree;
vector<lower=0,upper=1>[Noent] z_aug; //augmented ordinal data
vector[len_free_c[1]] Lambda_y_free_c;
vector[len_free_c[4]] B_free_c;
vector<lower=0>[len_free_c[5]] Theta_sd_free_c;
vector<lower=-1,upper=1>[len_free_c[7]] Theta_r_free_c; // to use beta prior
vector<lower=0>[len_free_c[9]] Psi_sd_free_c;
array[nblk_c[1]] corr_matrix[psidims_c[1]] Psi_r_mat_1_c;
array[nblk_c[2]] corr_matrix[psidims_c[2]] Psi_r_mat_2_c;
array[nblk_c[3]] corr_matrix[psidims_c[3]] Psi_r_mat_3_c;
array[nblk_c[4]] corr_matrix[psidims_c[4]] Psi_r_mat_4_c;
array[nblk_c[5]] corr_matrix[psidims_c[5]] Psi_r_mat_5_c;
vector<lower=-1,upper=1>[len_free_c[10]] Psi_r_free_c;
vector[len_free_c[13]] Nu_free_c;
vector[len_free_c[14]] Alpha_free_c;
}
transformed parameters {
array[Ng] matrix[p, m] Lambda_y;
array[Ng] matrix[m, m] B;
array[Ng] matrix[p, p] Theta_sd;
array[Ng] matrix[p, p] T_r_lower;
array[Ng] matrix[p, p] Theta_r;
array[Ng] matrix[p + q, 1] Nu;
array[Ng] matrix[m + n, 1] Alpha;
array[Ng] matrix[p_c, m_c] Lambda_y_c;
array[Ng] matrix[m_c, m_c] B_c;
array[Ng] matrix[p_c, p_c] Theta_sd_c;
array[Ng] matrix[p_c, p_c] T_r_lower_c;
array[Ng] matrix[p_c, p_c] Theta_r_c;
array[Ng] matrix[p_c, 1] Nu_c;
array[Ng] matrix[m_c, 1] Alpha_c;
array[Ng] matrix[sum(nlevs) - Nord, 1] Tau_un;
array[Ng] matrix[sum(nlevs) - Nord, 1] Tau;
vector[len_free[15]] Tau_free;
real tau_jacobian;
array[Ng] matrix[m, m] Psi;
array[Ng] matrix[m, m] Psi_sd;
array[Ng] matrix[m, m] Psi_r_lower;
array[Ng] matrix[m, m] Psi_r;
array[Ng] matrix[m_c, m_c] Psi_c;
array[Ng] matrix[m_c, m_c] Psi_sd_c;
array[Ng] matrix[m_c, m_c] Psi_r_lower_c;
array[Ng] matrix[m_c, m_c] Psi_r_c;
vector[len_free[1]] lambda_y_primn;
vector[len_free[4]] b_primn;
vector[len_free[13]] nu_primn;
vector[len_free[14]] alpha_primn;
vector[len_free[15]] tau_primn;
vector[len_free_c[1]] lambda_y_primn_c;
vector[len_free_c[4]] b_primn_c;
vector[len_free_c[13]] nu_primn_c;
vector[len_free_c[14]] alpha_primn_c;
array[Ng] matrix[p, m] Lambda_y_A; // = Lambda_y * (I - B)^{-1}
array[Ng] matrix[p_c, m_c] Lambda_y_A_c;
array[Ng] vector[p + q] Mu;
array[Ng] matrix[p + q, p + q] Sigma; // model covariance matrix
array[Ng] matrix[p + q, p + q] Sigmainv_grp; // model covariance matrix
array[Ng] real logdetSigma_grp;
array[Np] matrix[p + q + 1, p + q + 1] Sigmainv; // for updating S^-1 by missing data pattern
array[Ng] vector[p_c] Mu_c;
array[Ng] matrix[p_c, p_c] Sigma_c; // level 2 model covariance matrix
array[Ng] matrix[N_both + N_within, N_both + N_within] S_PW;
array[Ntot] vector[p + q] YXstar;
array[Ntot] vector[Nord] YXostar; // ordinal data
for (g in 1:Ng) {
// model matrices
Lambda_y[g] = fill_matrix(Lambda_y_free, Lambda_y_skeleton[g], w1skel, g_start1[g,1], g_start1[g,2]);
B[g] = fill_matrix(B_free, B_skeleton[g], w4skel, g_start4[g,1], g_start4[g,2]);
Theta_sd[g] = fill_matrix(Theta_sd_free, Theta_skeleton[g], w5skel, g_start5[g,1], g_start5[g,2]);
T_r_lower[g] = fill_matrix(Theta_r_free, Theta_r_skeleton[g], w7skel, g_start7[g,1], g_start7[g,2]);
Theta_r[g] = T_r_lower[g] + transpose(T_r_lower[g]) - diag_matrix(rep_vector(1, p));
if (!use_cov) {
Nu[g] = fill_matrix(Nu_free, Nu_skeleton[g], w13skel, g_start13[g,1], g_start13[g,2]);
Alpha[g] = fill_matrix(Alpha_free, Alpha_skeleton[g], w14skel, g_start14[g,1], g_start14[g,2]);
}
Psi[g] = diag_matrix(rep_vector(0, m));
if (m > 0) {
Psi_sd[g] = fill_matrix(Psi_sd_free, Psi_skeleton[g], w9skel, g_start9[g,1], g_start9[g,2]);
Psi_r_lower[g] = fill_matrix(Psi_r_free, Psi_r_skeleton[g], w10skel, g_start10[g,1], g_start10[g,2]);
Psi_r[g] = Psi_r_lower[g] + transpose(Psi_r_lower[g]) - diag_matrix(rep_vector(1, m));
}
// level 2 matrices
Lambda_y_c[g] = fill_matrix(Lambda_y_free_c, Lambda_y_skeleton_c[g], w1skel_c, g_start1_c[g,1], g_start1_c[g,2]);
B_c[g] = fill_matrix(B_free_c, B_skeleton_c[g], w4skel_c, g_start4_c[g,1], g_start4_c[g,2]);
Theta_sd_c[g] = fill_matrix(Theta_sd_free_c, Theta_skeleton_c[g], w5skel_c, g_start5_c[g,1], g_start5_c[g,2]);
T_r_lower_c[g] = fill_matrix(Theta_r_free_c, Theta_r_skeleton_c[g], w7skel_c, g_start7_c[g,1], g_start7_c[g,2]);
Theta_r_c[g] = T_r_lower_c[g] + transpose(T_r_lower_c[g]) - diag_matrix(rep_vector(1, p_c));
Nu_c[g] = fill_matrix(Nu_free_c, Nu_skeleton_c[g], w13skel_c, g_start13_c[g,1], g_start13_c[g,2]);
Alpha_c[g] = fill_matrix(Alpha_free_c, Alpha_skeleton_c[g], w14skel_c, g_start14_c[g,1], g_start14_c[g,2]);
Psi_c[g] = diag_matrix(rep_vector(0, m_c));
if (m_c > 0) {
Psi_sd_c[g] = fill_matrix(Psi_sd_free_c, Psi_skeleton_c[g], w9skel_c, g_start9_c[g,1], g_start9_c[g,2]);
Psi_r_lower_c[g] = fill_matrix(Psi_r_free_c, Psi_r_skeleton_c[g], w10skel_c, g_start10_c[g,1], g_start10_c[g,2]);
Psi_r_c[g] = Psi_r_lower_c[g] + transpose(Psi_r_lower_c[g]) - diag_matrix(rep_vector(1, m_c));
}
}
if (sum(nblk) > 0) {
// we need to define a separate parameter for each dimension of correlation matrix,
// so we need all these Psi_r_mats
Psi_r = fill_cov(Psi_r, blkse, nblk, Psi_r_mat_1, Psi_r_mat_2, Psi_r_mat_3, Psi_r_mat_4, Psi_r_mat_5);
}
if (sum(nblk_c) > 0) {
Psi_r_c = fill_cov(Psi_r_c, blkse_c, nblk_c, Psi_r_mat_1_c, Psi_r_mat_2_c, Psi_r_mat_3_c, Psi_r_mat_4_c, Psi_r_mat_5_c);
}
// see https://books.google.com/books?id=9AC-s50RjacC&lpg=PP1&dq=LISREL&pg=PA3#v=onepage&q=LISREL&f=false
for (g in 1:Ng) {
if (m > 0) {
Lambda_y_A[g] = mdivide_right(Lambda_y[g], I - B[g]); // = Lambda_y * (I - B)^{-1}
Psi[g] = quad_form_sym(Psi_r[g], Psi_sd[g]);
}
if (!use_cov) {
Mu[g] = to_vector(Nu[g]);
} else if(has_data) {
Mu[g] = YXbar[g]; // doesn't enter in likelihood, just for lppd + loo
}
if (p > 0) {
Sigma[g, 1:p, 1:p] = quad_form_sym(Theta_r[g], Theta_sd[g]);
if (m > 0) {
Sigma[g, 1:p, 1:p] += quad_form_sym(Psi[g], transpose(Lambda_y_A[g]));
if (!use_cov) Mu[g, 1:p] += to_vector(Lambda_y_A[g] * Alpha[g, 1:m, 1]);
}
}
if (m_c > 0) {
Lambda_y_A_c[g] = mdivide_right(Lambda_y_c[g], I_c - B_c[g]);
Psi_c[g] = quad_form_sym(Psi_r_c[g], Psi_sd_c[g]);
}
Mu_c[g] = to_vector(Nu_c[g]);
if (p_c > 0) {
Sigma_c[g, 1:p_c, 1:p_c] = quad_form_sym(Theta_r_c[g], Theta_sd_c[g]);
if (m_c > 0) {
Sigma_c[g, 1:p_c, 1:p_c] += quad_form_sym(Psi_c[g], transpose(Lambda_y_A_c[g]));
Mu_c[g, 1:p_c] += to_vector(Lambda_y_A_c[g] * Alpha_c[g, 1:m_c, 1]);
}
}
if (nclus[g,2] > 1) {
// remove between variables, for likelihood computations
S_PW[g] = cov_w[g, between_idx[(N_between + 1):p_tilde], between_idx[(N_between + 1):p_tilde]];
}
}
// obtain ordered thresholds; NB untouched for two-level models
if (ord) {
int opos = 1;
int ofreepos = 1;
tau_jacobian = 0;
for (g in 1:Ng) {
int vecpos = 1;
Tau_un[g] = fill_matrix(Tau_ufree, Tau_skeleton[g], w15skel, g_start15[g,1], g_start15[g,2]);
for (i in 1:Nord) {
for (j in 1:(nlevs[i] - 1)) {
real rc = Tau_skeleton[g, vecpos, 1];
int eq = w15skel[opos, 1];
int wig = w15skel[opos, 3];
if (is_inf(rc)) {
if (eq == 0 || wig == 1) {
if (j == 1) {
Tau[g, vecpos, 1] = Tau_un[g, vecpos, 1];
} else {
Tau[g, vecpos, 1] = Tau[g, (vecpos - 1), 1] + exp(Tau_un[g, vecpos, 1]);
}
Tau_free[ofreepos] = Tau[g, vecpos, 1];
// this is used if a prior goes on Tau_free, instead of Tau_ufree:
//if (j > 1) {
// tau_jacobian += Tau_un[g, vecpos, 1]; // see https://mc-stan.org/docs/2_24/reference-manual/ordered-vector.html
// }
ofreepos += 1;
} else if (eq == 1) {
int eqent = w15skel[opos, 2];
Tau[g, vecpos, 1] = Tau_free[eqent];
}
opos += 1;
} else {
// fixed value
Tau[g, vecpos, 1] = Tau_un[g, vecpos, 1];
}
vecpos += 1;
}
}
}
}
// prior vectors
if (wigind) {
lambda_y_primn = fill_prior(Lambda_y_free, lambda_y_mn, w1skel);
b_primn = fill_prior(B_free, b_mn, w4skel);
nu_primn = fill_prior(Nu_free, nu_mn, w13skel);
alpha_primn = fill_prior(Alpha_free, alpha_mn, w14skel);
tau_primn = fill_prior(Tau_ufree, tau_mn, w15skel);
lambda_y_primn_c = fill_prior(Lambda_y_free_c, lambda_y_mn_c, w1skel_c);
b_primn_c = fill_prior(B_free_c, b_mn_c, w4skel_c);
nu_primn_c = fill_prior(Nu_free_c, nu_mn_c, w13skel_c);
alpha_primn_c = fill_prior(Alpha_free_c, alpha_mn_c, w14skel_c);
} else {
lambda_y_primn = to_vector(lambda_y_mn);
b_primn = to_vector(b_mn);
nu_primn = to_vector(nu_mn);
alpha_primn = to_vector(alpha_mn);
tau_primn = to_vector(tau_mn);
lambda_y_primn_c = to_vector(lambda_y_mn_c);
b_primn_c = to_vector(b_mn_c);
nu_primn_c = to_vector(nu_mn_c);
alpha_primn_c = to_vector(alpha_mn_c);
}
// NB nothing below this will be used for two level, because we need other tricks to
// compute the likelihood
// continuous responses underlying ordinal data
if (ord) {
int idxvec = 0;
for (patt in 1:Np) {
for (i in startrow[patt]:endrow[patt]) {
for (j in 1:Nordobs[patt]) {
int obspos = OrdObsvar[patt,j];
int vecpos = YXo[i,obspos] - 1;
idxvec += 1;
if (obspos > 1) vecpos += sum(nlevs[1:(obspos - 1)]) - (obspos - 1);
if (YXo[i,obspos] == 1) {
YXostar[i,obspos] = -10 + (Tau[grpnum[patt], (vecpos + 1), 1] + 10) .* z_aug[idxvec];
tau_jacobian += log(abs(Tau[grpnum[patt], (vecpos + 1), 1] + 10)); // must add log(U) to tau_jacobian
} else if (YXo[i,obspos] == nlevs[obspos]) {
YXostar[i,obspos] = Tau[grpnum[patt], vecpos, 1] + (10 - Tau[grpnum[patt], vecpos, 1]) .* z_aug[idxvec];
tau_jacobian += log(abs(10 - Tau[grpnum[patt], vecpos, 1]));
} else {
YXostar[i,obspos] = Tau[grpnum[patt], vecpos, 1] + (Tau[grpnum[patt], (vecpos + 1), 1] - Tau[grpnum[patt], vecpos, 1]) .* z_aug[idxvec];
tau_jacobian += Tau_un[grpnum[patt], (vecpos + 1), 1]; // jacobian is log(exp(Tau_un))
}
YXstar[i, ordidx[obspos]] = YXostar[i, obspos];
}
}
}
}
if (Ncont > 0) {
for (patt in 1:Np) {
for (i in startrow[patt]:endrow[patt]) {
for (j in 1:Ncont) {
YXstar[i, contidx[j]] = YX[i,j];
}
}
}
}
// move observations to the left
if (missing) {
for (patt in 1:Np) {
for (i in startrow[patt]:endrow[patt]) {
for (j in 1:Nobs[patt]) {
YXstar[i,j] = YXstar[i, Obsvar[patt,j]];
}
}
}
}
// for computing mvn with sufficient stats
if (!multilev) {
for (g in 1:Ng) {
Sigmainv_grp[g] = inverse_spd(Sigma[g]);
logdetSigma_grp[g] = log_determinant(Sigma[g]);
}
for (patt in 1:Np) {
Sigmainv[patt, 1:(Nobs[patt] + 1), 1:(Nobs[patt] + 1)] = sig_inv_update(Sigmainv_grp[grpnum[patt]], Obsvar[patt,], Nobs[patt], p + q, logdetSigma_grp[grpnum[patt]]);
}
}
}
model { // N.B.: things declared in the model block do not get saved in the output, which is okay here
/* transformed sd parameters for priors */
vector[len_free[5]] Theta_pri;
vector[len_free[9]] Psi_pri;
vector[len_free_c[5]] Theta_pri_c;
vector[len_free_c[9]] Psi_pri_c;
/* log-likelihood */
if (multilev && has_data) {
int grpidx;
int r1 = 1; // index clusters per group
int r2 = 0;
int rr1 = 1; // index units per group
int rr2 = 0;
int r3 = 1; // index unique cluster sizes per group
int r4 = 0;
for (mm in 1:Np) {
grpidx = grpnum[mm];
if (grpidx > 1) {
r1 += nclus[(grpidx - 1), 2];
rr1 += nclus[(grpidx - 1), 1];
r3 += ncluster_sizes[(grpidx - 1)];
}
r2 += nclus[grpidx, 2];
rr2 += nclus[grpidx, 1];
r4 += ncluster_sizes[grpidx];
target += twolevel_logdens(mean_d[r3:r4], cov_d[r3:r4], S_PW[grpidx], YX[rr1:rr2],
nclus[grpidx,], cluster_size[r1:r2], cluster_sizes[r3:r4],
ncluster_sizes[grpidx], cluster_size_ns[r3:r4], Mu[grpidx],
Sigma[grpidx], Mu_c[grpidx], Sigma_c[grpidx],
ov_idx1, ov_idx2, within_idx, between_idx, both_idx,
p_tilde, N_within, N_between, N_both);
if (Nx[grpidx] + Nx_between[grpidx] > 0) target += -log_lik_x;
}
} else if (use_cov && !pri_only) {
for (g in 1:Ng) {
target += wishart_lpdf((N[g] - 1) * Sstar[g] | N[g] - 1, Sigma[g]);
if (Nx[g] > 0) {
array[Nx[g]] int xvars = Xdatvar[g, 1:Nx[g]];
target += -wishart_lpdf((N[g] - 1) * Sstar[g, xvars, xvars] | N[g] - 1, Sigma[g, xvars, xvars]);
}
}
} else if (has_data && !pri_only) {
array[p + q] int obsidx;
array[p + q] int xidx;
array[p + q] int xdatidx;
int grpidx;
int r1;
int r2;
for (mm in 1:Np) {
obsidx = Obsvar[mm,];
xidx = Xvar[mm,];
xdatidx = Xdatvar[mm,];
grpidx = grpnum[mm];
r1 = startrow[mm];
r2 = endrow[mm];
if (!use_suff) {
target += multi_normal_lpdf(YXstar[r1:r2,1:Nobs[mm]] | Mu[grpidx, obsidx[1:Nobs[mm]]], Sigma[grpidx, obsidx[1:Nobs[mm]], obsidx[1:Nobs[mm]]]);
if (Nx[mm] > 0) {
target += -multi_normal_lpdf(YXstar[r1:r2,xdatidx[1:Nx[mm]]] | Mu[grpidx, xidx[1:Nx[mm]]], Sigma[grpidx, xidx[1:Nx[mm]], xidx[1:Nx[mm]]]);
}
} else {
// sufficient stats
target += multi_normal_suff(YXbarstar[mm, 1:Nobs[mm]], Sstar[mm, 1:Nobs[mm], 1:Nobs[mm]], Mu[grpidx, obsidx[1:Nobs[mm]]], Sigmainv[mm, 1:(Nobs[mm] + 1), 1:(Nobs[mm] + 1)], r2 - r1 + 1);
if (Nx[mm] > 0) {
target += -multi_normal_suff(YXbarstar[mm, xdatidx[1:Nx[mm]]], Sstar[mm, xdatidx[1:Nx[mm]], xdatidx[1:Nx[mm]]], Mu[grpidx, xidx[1:Nx[mm]]], sig_inv_update(Sigmainv[grpidx], xidx, Nx[mm], p + q, logdetSigma_grp[grpidx]), r2 - r1 + 1);
}
}
}
if (ord) {
target += tau_jacobian;
}
}
/* prior densities in log-units */
target += normal_lpdf(Lambda_y_free | lambda_y_primn, lambda_y_sd);
target += normal_lpdf(B_free | b_primn, b_sd);
target += normal_lpdf(Nu_free | nu_primn, nu_sd);
target += normal_lpdf(Alpha_free | alpha_primn, alpha_sd);
target += normal_lpdf(Tau_ufree | tau_primn, tau_sd);
target += normal_lpdf(Lambda_y_free_c | lambda_y_primn_c, lambda_y_sd_c);
target += normal_lpdf(B_free_c | b_primn_c, b_sd_c);
target += normal_lpdf(Nu_free_c | nu_primn_c, nu_sd_c);
target += normal_lpdf(Alpha_free_c | alpha_primn_c, alpha_sd_c);
/* transform sd parameters to var or prec, depending on
what the user wants. */
Theta_pri = Theta_sd_free;
if (len_free[5] > 0 && theta_pow != 1) {
for (i in 1:len_free[5]) {
Theta_pri[i] = Theta_sd_free[i]^(theta_pow);
target += log(abs(theta_pow)) + (theta_pow - 1)*log(Theta_sd_free[i]);
}
}
Psi_pri = Psi_sd_free;
if (len_free[9] > 0 && psi_pow != 1) {
for (i in 1:len_free[9]) {
Psi_pri[i] = Psi_sd_free[i]^(psi_pow);
target += log(abs(psi_pow)) + (psi_pow - 1)*log(Psi_sd_free[i]);
}
}
target += gamma_lpdf(Theta_pri | theta_sd_shape, theta_sd_rate);
target += gamma_lpdf(Psi_pri | psi_sd_shape, psi_sd_rate);
target += beta_lpdf(.5 * (1 + Theta_r_free) | theta_r_alpha, theta_r_beta) + log(.5) * len_free[7]; // the latter term is the jacobian moving from (-1,1) to (0,1), because beta_lpdf is defined on (0,1)
if (sum(nblk) > 0) {
for (k in 1:sum(nblk)) {
int blkidx = blkse[k, 6];
int arrayidx = blkse[k, 5];
if (arrayidx == 1) {
target += lkj_corr_lpdf(Psi_r_mat_1[blkidx] | blkse[k,7]);
} else if (arrayidx == 2) {
target += lkj_corr_lpdf(Psi_r_mat_2[blkidx] | blkse[k,7]);
} else if (arrayidx == 3) {
target += lkj_corr_lpdf(Psi_r_mat_3[blkidx] | blkse[k,7]);
} else if (arrayidx == 4) {
target += lkj_corr_lpdf(Psi_r_mat_4[blkidx] | blkse[k,7]);
} else {
target += lkj_corr_lpdf(Psi_r_mat_5[blkidx] | blkse[k,7]);
}
}
}
if (len_free[10] > 0) {
target += beta_lpdf(.5 * (1 + Psi_r_free) | psi_r_alpha, psi_r_beta) + log(.5) * len_free[10];
}
// and the same for level 2
Theta_pri_c = Theta_sd_free_c;
if (len_free_c[5] > 0 && theta_pow_c != 1) {
for (i in 1:len_free_c[5]) {
Theta_pri_c[i] = Theta_sd_free_c[i]^(theta_pow_c);
target += log(abs(theta_pow_c)) + (theta_pow_c - 1)*log(Theta_sd_free_c[i]);
}
}
Psi_pri_c = Psi_sd_free_c;
if (len_free_c[9] > 0 && psi_pow_c != 1) {
for (i in 1:len_free_c[9]) {
Psi_pri_c[i] = Psi_sd_free_c[i]^(psi_pow_c);
target += log(abs(psi_pow_c)) + (psi_pow_c - 1)*log(Psi_sd_free_c[i]);
}
}
target += gamma_lpdf(Theta_pri_c | theta_sd_shape_c, theta_sd_rate_c);
target += gamma_lpdf(Psi_pri_c | psi_sd_shape_c, psi_sd_rate_c);
target += beta_lpdf(.5 * (1 + Theta_r_free_c) | theta_r_alpha_c, theta_r_beta_c) + log(.5) * len_free_c[7];
if (sum(nblk_c) > 0) {
for (k in 1:sum(nblk_c)) {
int blkidx = blkse_c[k, 6];
int arrayidx = blkse_c[k, 5];
if (arrayidx == 1) {
target += lkj_corr_lpdf(Psi_r_mat_1_c[blkidx] | blkse_c[k,7]);
} else if (arrayidx == 2) {
target += lkj_corr_lpdf(Psi_r_mat_2_c[blkidx] | blkse_c[k,7]);
} else if (arrayidx == 3) {
target += lkj_corr_lpdf(Psi_r_mat_3_c[blkidx] | blkse_c[k,7]);
} else if (arrayidx == 4) {
target += lkj_corr_lpdf(Psi_r_mat_4_c[blkidx] | blkse_c[k,7]);
} else {
target += lkj_corr_lpdf(Psi_r_mat_5_c[blkidx] | blkse_c[k,7]);
}
}
} else if (len_free_c[10] > 0) {
target += beta_lpdf(.5 * (1 + Psi_r_free_c) | psi_r_alpha_c, psi_r_beta_c) + log(.5) * len_free_c[10];
}
}
generated quantities { // these matrices are saved in the output but do not figure into the likelihood
// see https://books.google.com/books?id=9AC-s50RjacC&lpg=PP1&dq=LISREL&pg=PA34#v=onepage&q=LISREL&f=false
// sign constraints and correlations
vector[len_free[1]] ly_sign;
vector[len_free[4]] bet_sign;
array[Ng] matrix[m, m] PSmat;
array[Ng] matrix[m, m] PS;
vector[len_free[7]] Theta_cov;
vector[len_free[5]] Theta_var;
vector[len_free[10]] P_r;
vector[len_free[11]] Psi_cov;
vector[len_free[9]] Psi_var;
// level 2
vector[len_free_c[1]] ly_sign_c;
vector[len_free_c[4]] bet_sign_c;
array[Ng] matrix[m_c, m_c] PSmat_c;
array[Ng] matrix[m_c, m_c] PS_c;
vector[len_free_c[7]] Theta_cov_c;
vector[len_free_c[5]] Theta_var_c;
vector[len_free_c[10]] P_r_c;
vector[len_free_c[11]] Psi_cov_c;
vector[len_free_c[9]] Psi_var_c;
// loglik + ppp
vector[multilev ? sum(nclus[,2]) : (use_cov ? Ng : Ntot)] log_lik; // for loo, etc
vector[multilev ? sum(nclus[,2]) : (use_cov ? Ng : Ntot)] log_lik_sat; // for ppp
array[Ntot] vector[multilev ? p_tilde : p + q] YXstar_rep; // artificial data
vector[multilev ? sum(nclus[,2]) : (use_cov ? Ng : Ntot)] log_lik_rep; // for loo, etc
vector[multilev ? sum(nclus[,2]) : (use_cov ? Ng : Ntot)] log_lik_rep_sat; // for ppp
array[Ng] matrix[p + q, p + q + 1] satout;
array[Ng] matrix[p + q, p + q + 1] satrep_out;
array[Ng] vector[p + q] Mu_sat;
array[Ng] matrix[p + q, p + q] Sigma_sat;
array[Ng] matrix[p + q, p + q] Sigma_sat_inv_grp;
array[Ng] real logdetS_sat_grp;
array[Np] matrix[p + q + 1, p + q + 1] Sigma_sat_inv;
array[Ng] vector[p + q] Mu_rep_sat;
array[Ng] matrix[p + q, p + q] Sigma_rep_sat;
array[Ng] matrix[p + q, p + q] Sigma_rep_sat_inv_grp;
array[Np] matrix[p + q + 1, p + q + 1] Sigma_rep_sat_inv;
array[Ng] real logdetS_rep_sat_grp;
matrix[p + q, p + q] zmat;
array[sum(nclus[,2])] vector[p_tilde] mean_d_rep;
vector[multilev ? sum(nclus[,2]) : Ng] log_lik_x_rep;
array[Ng] matrix[N_both + N_within, N_both + N_within] S_PW_rep;
array[Ng] matrix[p_tilde, p_tilde] S_PW_rep_full;
array[Ng] vector[p_tilde] ov_mean_rep;
array[Ng] vector[p_tilde] xbar_b_rep;
array[Ng] matrix[N_between, N_between] S2_rep;
array[Ng] matrix[p_tilde, p_tilde] S_B_rep;
array[Ng] matrix[p_tilde, p_tilde] cov_b_rep;
real<lower=0, upper=1> ppp;
// first deal with sign constraints:
ly_sign = sign_constrain_load(Lambda_y_free, len_free[1], lam_y_sign);
bet_sign = sign_constrain_reg(B_free, len_free[4], b_sign, Lambda_y_free, Lambda_y_free);
if (len_free[10] > 0) {
P_r = sign_constrain_reg(Psi_r_free, len_free[10], psi_r_sign, Lambda_y_free, Lambda_y_free);
}
ly_sign_c = sign_constrain_load(Lambda_y_free_c, len_free_c[1], lam_y_sign_c);
bet_sign_c = sign_constrain_reg(B_free_c, len_free_c[4], b_sign_c, Lambda_y_free_c, Lambda_y_free_c);
if (len_free_c[10] > 0) {
P_r_c = sign_constrain_reg(Psi_r_free_c, len_free_c[10], psi_r_sign_c, Lambda_y_free_c, Lambda_y_free_c);
}
for (g in 1:Ng) {
if (m > 0) {
PSmat[g] = fill_matrix(P_r, Psi_r_skeleton[g], w10skel, g_start10[g,1], g_start10[g,2]) + transpose(fill_matrix(P_r, Psi_r_skeleton[g], w10skel, g_start10[g,1], g_start10[g,2])) - diag_matrix(rep_vector(1, m));
}
if (m_c > 0) {
PSmat_c[g] = fill_matrix(P_r_c, Psi_r_skeleton_c[g], w10skel_c, g_start10_c[g,1], g_start10_c[g,2]) + transpose(fill_matrix(P_r_c, Psi_r_skeleton_c[g], w10skel_c, g_start10_c[g,1], g_start10_c[g,2])) - diag_matrix(rep_vector(1, m_c));
}
}
if (sum(nblk) > 0) {
PSmat = fill_cov(PSmat, blkse, nblk, Psi_r_mat_1, Psi_r_mat_2, Psi_r_mat_3, Psi_r_mat_4, Psi_r_mat_5);
}
if (sum(nblk_c) > 0) {
PSmat_c = fill_cov(PSmat_c, blkse_c, nblk_c, Psi_r_mat_1_c, Psi_r_mat_2_c, Psi_r_mat_3_c, Psi_r_mat_4_c, Psi_r_mat_5_c);
}
for (g in 1:Ng) {
PS[g] = quad_form_sym(PSmat[g], Psi_sd[g]);
PS_c[g] = quad_form_sym(PSmat_c[g], Psi_sd_c[g]);
}
// off-diagonal covariance parameter vectors, from cor/sd matrices:
Theta_cov = cor2cov(Theta_r, Theta_sd, num_elements(Theta_r_free), Theta_r_skeleton, w7skel, Ng);
Theta_var = Theta_sd_free .* Theta_sd_free;
if (m > 0 && len_free[11] > 0) {
/* iden is created so that we can re-use cor2cov, even though
we don't need to multiply to get covariances */
array[Ng] matrix[m, m] iden;
for (g in 1:Ng) {
iden[g] = diag_matrix(rep_vector(1, m));
}
Psi_cov = cor2cov(PS, iden, len_free[11], Psi_r_skeleton_f, w11skel, Ng);
} else {
Psi_cov = P_r;
}
Psi_var = Psi_sd_free .* Psi_sd_free;
// and for level 2
Theta_cov_c = cor2cov(Theta_r_c, Theta_sd_c, num_elements(Theta_r_free_c), Theta_r_skeleton_c, w7skel_c, Ng);
Theta_var_c = Theta_sd_free_c .* Theta_sd_free_c;
if (m_c > 0 && len_free_c[11] > 0) {
array[Ng] matrix[m_c, m_c] iden_c;
for (g in 1:Ng) {
iden_c[g] = diag_matrix(rep_vector(1, m_c));
}
Psi_cov_c = cor2cov(PS_c, iden_c, len_free_c[11], Psi_r_skeleton_f_c, w11skel_c, Ng);
} else {
Psi_cov_c = P_r_c;
}
Psi_var_c = Psi_sd_free_c .* Psi_sd_free_c;
{ // log-likelihood
array[p + q] int obsidx;
array[p + q] int xidx;
array[p + q] int xdatidx;
int r1;
int r2;
int r3;
int r4;
int rr1;
int rr2;
int grpidx;
int clusidx;
if (do_test && use_cov) {
for (g in 1:Ng) {
Sigma_rep_sat[g] = wishart_rng(N[g] - 1, Sigma[g]);
}
} else if (do_test && has_data) {
// generate level 2 data, then level 1
if (multilev) {
array[p_tilde - N_between] int notbidx;
notbidx = between_idx[(N_between + 1):p_tilde];
r1 = 1;
rr1 = 1;
clusidx = 1;
r2 = 1;
for (gg in 1:Ng) {
matrix[p_c, p_c] Sigma_c_chol = cholesky_decompose(Sigma_c[gg]);
matrix[p + q, p + q] Sigma_chol = cholesky_decompose(Sigma[gg]);
S_PW_rep[gg] = rep_matrix(0, N_both + N_within, N_both + N_within);
S_PW_rep_full[gg] = rep_matrix(0, p_tilde, p_tilde);
S_B_rep[gg] = rep_matrix(0, p_tilde, p_tilde);
ov_mean_rep[gg] = rep_vector(0, p_tilde);
for (cc in 1:nclus[gg, 2]) {
vector[p_c] YXstar_rep_c;
vector[p_tilde] YXstar_rep_tilde;
YXstar_rep_c = multi_normal_cholesky_rng(Mu_c[gg], Sigma_c_chol);
YXstar_rep_tilde = calc_B_tilde(Sigma_c[gg], YXstar_rep_c, ov_idx2, p_tilde)[,1];
for (ii in r1:(r1 + cluster_size[clusidx] - 1)) {
vector[N_within + N_both] Ywb_rep;
Ywb_rep = multi_normal_cholesky_rng(Mu[gg], Sigma_chol);
YXstar_rep[ii] = YXstar_rep_tilde;
for (ww in 1:(p_tilde - N_between)) {
YXstar_rep[ii, notbidx[ww]] += Ywb_rep[ww];
}
ov_mean_rep[gg] += YXstar_rep[ii];
}
for (jj in 1:p_tilde) {
mean_d_rep[clusidx, jj] = mean(YXstar_rep[r1:(r1 + cluster_size[clusidx] - 1), jj]);
}
r1 += cluster_size[clusidx];
clusidx += 1;
} // cc
ov_mean_rep[gg] *= pow(nclus[gg, 1], -1);
xbar_b_rep[gg] = ov_mean_rep[gg];
r1 -= nclus[gg, 1]; // reset for S_PW
clusidx -= nclus[gg, 2];
if (N_between > 0) {
S2_rep[gg] = rep_matrix(0, N_between, N_between);
for (ii in 1:N_between) {
xbar_b_rep[gg, between_idx[ii]] = mean(mean_d_rep[clusidx:(clusidx + nclus[gg, 2] - 1), between_idx[ii]]);
}
}
for (cc in 1:nclus[gg, 2]) {
for (ii in r1:(r1 + cluster_size[clusidx] - 1)) {
S_PW_rep_full[gg] += tcrossprod(to_matrix(YXstar_rep[ii] - mean_d_rep[clusidx]));
}
S_B_rep[gg] += cluster_size[clusidx] * tcrossprod(to_matrix(mean_d_rep[clusidx] - ov_mean_rep[gg]));
if (N_between > 0) {
S2_rep[gg] += tcrossprod(to_matrix(mean_d_rep[clusidx, between_idx[1:N_between]] - xbar_b_rep[gg, between_idx[1:N_between]]));
}
r1 += cluster_size[clusidx];
clusidx += 1;
}
S_PW_rep_full[gg] *= pow(nclus[gg, 1] - nclus[gg, 2], -1);
S_B_rep[gg] *= pow(nclus[gg, 2] - 1, -1);
S2_rep[gg] *= pow(nclus[gg, 2], -1);
// mods to between-only variables:
if (N_between > 0) {
array[N_between] int betonly = between_idx[1:N_between];
S_PW_rep_full[gg, betonly, betonly] = rep_matrix(0, N_between, N_between);
// Y2: mean_d_rep; Y2c: mean_d_rep - ov_mean_rep
for (ii in 1:N_between) {
for (jj in 1:(N_both + N_within)) {
S_B_rep[gg, between_idx[ii], between_idx[(N_between + jj)]] *= (gs[gg] * nclus[gg, 2] * pow(nclus[gg, 1], -1));
S_B_rep[gg, between_idx[(N_between + jj)], between_idx[ii]] = S_B_rep[gg, between_idx[ii], between_idx[(N_between + jj)]];
}
}
S_B_rep[gg, betonly, betonly] = rep_matrix(0, N_between, N_between);
for (cc in 1:nclus[gg, 2]) {
S_B_rep[gg, betonly, betonly] += tcrossprod(to_matrix(mean_d_rep[cc, betonly] - ov_mean_rep[gg, betonly]));
}
S_B_rep[gg, betonly, betonly] *= gs[gg] * pow(nclus[gg, 2], -1);
}
cov_b_rep[gg] = pow(gs[gg], -1) * (S_B_rep[gg] - S_PW_rep_full[gg]);
if (N_between > 0) {
cov_b_rep[gg, between_idx[1:N_between], between_idx[1:N_between]] = S2_rep[gg];
}
rr1 = r1 - nclus[gg, 1];
r2 = clusidx - nclus[gg, 2];
Mu_rep_sat[gg] = rep_vector(0, N_within + N_both);
if (N_within > 0) {
for (j in 1:N_within) {
xbar_b_rep[gg, within_idx[j]] = 0;
Mu_rep_sat[gg, within_idx[j]] = ov_mean_rep[gg, within_idx[j]];
}
}
S_PW_rep[gg] = S_PW_rep_full[gg, notbidx, notbidx];
if (Nx[gg] > 0 || Nx_between[gg] > 0) {
array[2] vector[p_tilde] mnvecs;
array[3] matrix[p_tilde, p_tilde] covmats;
mnvecs = calc_mean_vecs(YXstar_rep[rr1:(r1 - 1)], mean_d_rep[r2:(clusidx - 1)], nclus[gg], Xvar[gg], Xbetvar[gg], Nx[gg], Nx_between[gg], p_tilde);
covmats = calc_cov_mats(YXstar_rep[rr1:(r1 - 1)], mean_d_rep[r2:(clusidx - 1)], mnvecs, nclus[gg], Xvar[gg], Xbetvar[gg], Nx[gg], Nx_between[gg], p_tilde);
log_lik_x_rep[r2:(clusidx - 1)] = calc_log_lik_x(mean_d_rep[r2:(clusidx - 1)],
mnvecs[2], covmats[1],
covmats[2], covmats[3],
nclus[gg], cluster_size[r2:(clusidx - 1)],
Xvar[gg], Xbetvar[gg], Nx[gg], Nx_between[gg]);
} // Nx[gg] > 0
} // gg
} else {
for (mm in 1:Np) {
matrix[Nobs[mm], Nobs[mm]] Sigma_chol;
obsidx = Obsvar[mm,];
xidx = Xvar[mm,];
xdatidx = Xdatvar[mm,];
grpidx = grpnum[mm];
r1 = startrow[mm];
r2 = endrow[mm];
Sigma_chol = cholesky_decompose(Sigma[ grpidx, obsidx[1:Nobs[mm]], obsidx[1:Nobs[mm]] ]);
for (jj in r1:r2) {
YXstar_rep[jj, 1:Nobs[mm]] = multi_normal_cholesky_rng(Mu[grpidx, obsidx[1:Nobs[mm]]], Sigma_chol);
}
}
if (missing) {
// start values for Mu and Sigma
for (g in 1:Ng) {
Mu_sat[g] = rep_vector(0, p + q);
Mu_rep_sat[g] = Mu_sat[g];
Sigma_sat[g] = diag_matrix(rep_vector(1, p + q));
Sigma_rep_sat[g] = Sigma_sat[g];
}
for (jj in 1:emiter) {
satout = estep(YXstar, Mu_sat, Sigma_sat, Nobs, Obsvar, startrow, endrow, grpnum, Np, Ng);
satrep_out = estep(YXstar_rep, Mu_rep_sat, Sigma_rep_sat, Nobs, Obsvar, startrow, endrow, grpnum, Np, Ng);
// M step
for (g in 1:Ng) {
Mu_sat[g] = satout[g,,1]/N[g];
Sigma_sat[g] = satout[g,,2:(p + q + 1)]/N[g] - Mu_sat[g] * Mu_sat[g]';
Mu_rep_sat[g] = satrep_out[g,,1]/N[g];
Sigma_rep_sat[g] = satrep_out[g,,2:(p + q + 1)]/N[g] - Mu_rep_sat[g] * Mu_rep_sat[g]';
}
}
} else {
// complete data; Np patterns must only correspond to groups
for (mm in 1:Np) {
array[3] int arr_dims = dims(YXstar);
matrix[endrow[mm] - startrow[mm] + 1, arr_dims[2]] YXsmat; // crossprod needs matrix
matrix[endrow[mm] - startrow[mm] + 1, arr_dims[2]] YXsrepmat;
r1 = startrow[mm];
r2 = endrow[mm];
grpidx = grpnum[mm];
for (jj in 1:(p + q)) {
Mu_sat[grpidx,jj] = mean(YXstar[r1:r2,jj]);
Mu_rep_sat[grpidx,jj] = mean(YXstar_rep[r1:r2,jj]);
}
for (jj in r1:r2) {
YXsmat[jj - r1 + 1] = (YXstar[jj] - Mu_sat[grpidx])';
YXsrepmat[jj - r1 + 1] = (YXstar_rep[jj] - Mu_rep_sat[grpidx])';
}
Sigma_sat[grpidx] = crossprod(YXsmat)/N[grpidx];
Sigma_rep_sat[grpidx] = crossprod(YXsrepmat)/N[grpidx];
// FIXME? Sigma_sat[grpidx] = tcrossprod(YXsmat); does not throw an error??
}
}
for (g in 1:Ng) {
Sigma_sat_inv_grp[g] = inverse_spd(Sigma_sat[g]);
logdetS_sat_grp[g] = log_determinant(Sigma_sat[g]);
Sigma_rep_sat_inv_grp[g] = inverse_spd(Sigma_rep_sat[g]);
logdetS_rep_sat_grp[g] = log_determinant(Sigma_rep_sat[g]);
}
for (mm in 1:Np) {
Sigma_sat_inv[mm, 1:(Nobs[mm] + 1), 1:(Nobs[mm] + 1)] = sig_inv_update(Sigma_sat_inv_grp[grpnum[mm]], Obsvar[mm,], Nobs[mm], p + q, logdetS_sat_grp[grpnum[mm]]);
Sigma_rep_sat_inv[mm, 1:(Nobs[mm] + 1), 1:(Nobs[mm] + 1)] = sig_inv_update(Sigma_rep_sat_inv_grp[grpnum[mm]], Obsvar[mm,], Nobs[mm], p + q, logdetS_rep_sat_grp[grpnum[mm]]);
}
}
}
// compute log-likelihoods
if (multilev) { // multilevel
r1 = 1;
r3 = 1;
r2 = 0;
r4 = 0;
for (mm in 1:Np) {
grpidx = grpnum[mm];
if (grpidx > 1) {
r1 += nclus[(grpidx - 1), 2];
r3 += nclus[(grpidx - 1), 1];
}
r2 += nclus[grpidx, 2];
r4 += nclus[grpidx, 1];
log_lik[r1:r2] = twolevel_logdens(mean_d_full[r1:r2], cov_d_full[r1:r2], S_PW[grpidx], YX[r3:r4],
nclus[grpidx,], cluster_size[r1:r2], cluster_size[r1:r2],
nclus[grpidx,2], intone[1:nclus[grpidx,2]], Mu[grpidx],
Sigma[grpidx], Mu_c[grpidx], Sigma_c[grpidx],
ov_idx1, ov_idx2, within_idx, between_idx, both_idx,
p_tilde, N_within, N_between, N_both);
if (Nx[grpidx] + Nx_between[grpidx] > 0) log_lik[r1:r2] -= log_lik_x_full[r1:r2];
}
}
zmat = rep_matrix(0, p + q, p + q);
// reset for 2-level loglik:
rr1 = 1;
r3 = 1;
rr2 = 0;
r4 = 0;
for (mm in 1:Np) {
obsidx = Obsvar[mm,];
xidx = Xvar[mm, 1:(p + q)];
xdatidx = Xdatvar[mm, 1:(p + q)];
grpidx = grpnum[mm];
r1 = startrow[mm];
r2 = endrow[mm];
if (use_cov) {
log_lik[mm] = wishart_lpdf((N[mm] - 1) * Sstar[mm] | N[mm] - 1, Sigma[mm]);
if (do_test) {
log_lik_sat[mm] = -log_lik[mm] + wishart_lpdf((N[mm] - 1) * Sstar[mm] | N[mm] - 1, Sstar[mm]);
log_lik_rep[mm] = wishart_lpdf(Sigma_rep_sat[mm] | N[mm] - 1, Sigma[mm]);
log_lik_rep_sat[mm] = wishart_lpdf(Sigma_rep_sat[mm] | N[mm] - 1, pow(N[mm] - 1, -1) * Sigma_rep_sat[mm]);
}
if (Nx[mm] > 0) {
array[Nx[mm]] int xvars = xdatidx[1:Nx[mm]];
log_lik[mm] += -wishart_lpdf((N[mm] - 1) * Sstar[mm, xvars, xvars] | N[mm] - 1, Sigma[mm, xvars, xvars]);
if (do_test) {
log_lik_sat[mm] += wishart_lpdf((N[mm] - 1) * Sstar[mm, xvars, xvars] | N[mm] - 1, Sigma[mm, xvars, xvars]);
log_lik_sat[mm] += -wishart_lpdf((N[mm] - 1) * Sstar[mm, xvars, xvars] | N[mm] - 1, Sstar[mm, xvars, xvars]);
log_lik_rep[mm] += -wishart_lpdf(Sigma_rep_sat[mm, xvars, xvars] | N[mm] - 1, Sigma[mm, xvars, xvars]);
log_lik_rep_sat[mm] += -wishart_lpdf(Sigma_rep_sat[mm, xvars, xvars] | N[mm] - 1, pow(N[mm] - 1, -1) * Sigma_rep_sat[mm, xvars, xvars]);
}
}
} else if (has_data && !multilev) {
for (jj in r1:r2) {
log_lik[jj] = multi_normal_suff(YXstar[jj, 1:Nobs[mm]], zmat[1:Nobs[mm], 1:Nobs[mm]], Mu[grpidx, obsidx[1:Nobs[mm]]], Sigmainv[mm], 1);
if (Nx[mm] > 0) {
log_lik[jj] += -multi_normal_suff(YXstar[jj, xdatidx[1:Nx[mm]]], zmat[1:Nx[mm], 1:Nx[mm]], Mu[grpidx, xidx[1:Nx[mm]]], sig_inv_update(Sigmainv[grpidx], xidx, Nx[mm], p + q, logdetSigma_grp[grpidx]), 1);
}
}
}
// saturated and y_rep likelihoods for ppp
if (do_test) {
if (multilev) {
// compute clusterwise log_lik_rep for grpidx
if (grpidx > 1) {
rr1 += nclus[(grpidx - 1), 2];
r3 += nclus[(grpidx - 1), 1];
}
rr2 += nclus[grpidx, 2];
r4 += nclus[grpidx, 1];
// NB: cov_d is 0 when we go cluster by cluster.
// otherwise it is covariance of cluster means by each unique cluster size
// because we go cluster by cluster here, we can reuse cov_d_full everywhere
log_lik_rep[rr1:rr2] = twolevel_logdens(mean_d_rep[rr1:rr2], cov_d_full[rr1:rr2],
S_PW_rep[grpidx], YXstar_rep[r3:r4],
nclus[grpidx,], cluster_size[rr1:rr2],
cluster_size[rr1:rr2], nclus[grpidx,2],
intone[1:nclus[grpidx,2]], Mu[grpidx],
Sigma[grpidx], Mu_c[grpidx], Sigma_c[grpidx],
ov_idx1, ov_idx2, within_idx, between_idx,
both_idx, p_tilde, N_within, N_between, N_both);
log_lik_sat[rr1:rr2] = twolevel_logdens(mean_d_full[rr1:rr2], cov_d_full[rr1:rr2],
S_PW[grpidx], YX[r3:r4],
nclus[grpidx,], cluster_size[rr1:rr2],
cluster_size[rr1:rr2], nclus[grpidx,2],
intone[1:nclus[grpidx,2]], xbar_w[grpidx, ov_idx1],
S_PW[grpidx], xbar_b[grpidx, ov_idx2], cov_b[grpidx, ov_idx2, ov_idx2],
ov_idx1, ov_idx2, within_idx, between_idx,
both_idx, p_tilde, N_within, N_between, N_both);
log_lik_rep_sat[rr1:rr2] = twolevel_logdens(mean_d_rep[rr1:rr2], cov_d_full[rr1:rr2],
S_PW_rep[grpidx], YXstar_rep[r3:r4],
nclus[grpidx,], cluster_size[rr1:rr2],
cluster_size[rr1:rr2], nclus[grpidx,2],
intone[1:nclus[grpidx,2]], Mu_rep_sat[grpidx],
S_PW_rep[grpidx], xbar_b_rep[grpidx, ov_idx2],
cov_b_rep[grpidx, ov_idx2, ov_idx2],
ov_idx1, ov_idx2,
within_idx, between_idx, both_idx, p_tilde,
N_within, N_between, N_both);
if (Nx[grpidx] + Nx_between[grpidx] > 0) {
log_lik_rep[rr1:rr2] -= log_lik_x_rep[rr1:rr2];
log_lik_sat[rr1:rr2] -= log_lik_x_full[rr1:rr2];
log_lik_rep_sat[rr1:rr2] -= log_lik_x_rep[rr1:rr2];
}
// we subtract log_lik here so that _sat always varies and does not lead to
// problems with rhat and neff computations
log_lik_sat[rr1:rr2] -= log_lik[rr1:rr2];
} else if (!use_cov) {
r1 = startrow[mm];
r2 = endrow[mm];
for (jj in r1:r2) {
log_lik_rep[jj] = multi_normal_suff(YXstar_rep[jj, 1:Nobs[mm]], zmat[1:Nobs[mm], 1:Nobs[mm]], Mu[grpidx, obsidx[1:Nobs[mm]]], Sigmainv[mm], 1);
log_lik_sat[jj] = multi_normal_suff(YXstar[jj, 1:Nobs[mm]], zmat[1:Nobs[mm], 1:Nobs[mm]], Mu_sat[grpidx, obsidx[1:Nobs[mm]]], Sigma_sat_inv[mm], 1);
log_lik_rep_sat[jj] = multi_normal_suff(YXstar_rep[jj, 1:Nobs[mm]], zmat[1:Nobs[mm], 1:Nobs[mm]], Mu_rep_sat[grpidx, obsidx[1:Nobs[mm]]], Sigma_rep_sat_inv[mm], 1);
// log_lik_sat, log_lik_sat_rep
if (Nx[mm] > 0) {
log_lik_rep[jj] += -multi_normal_suff(YXstar_rep[jj, xdatidx[1:Nx[mm]]], zmat[1:Nx[mm], 1:Nx[mm]], Mu[grpidx, xidx[1:Nx[mm]]], sig_inv_update(Sigmainv[grpidx], xidx, Nx[mm], p + q, logdetSigma_grp[grpidx]), 1);
log_lik_sat[jj] += -multi_normal_suff(YXstar[jj, xdatidx[1:Nx[mm]]], zmat[1:Nx[mm], 1:Nx[mm]], Mu_sat[grpidx, xidx[1:Nx[mm]]], sig_inv_update(Sigma_sat_inv[grpidx], xidx, Nx[mm], p + q, logdetS_sat_grp[grpidx]), 1);
log_lik_rep_sat[jj] += -multi_normal_suff(YXstar_rep[jj, xdatidx[1:Nx[mm]]], zmat[1:Nx[mm], 1:Nx[mm]], Mu_rep_sat[grpidx, xidx[1:Nx[mm]]], sig_inv_update(Sigma_rep_sat_inv[grpidx], xidx, Nx[mm], p + q, logdetS_rep_sat_grp[grpidx]), 1);
}
}
// we subtract log_lik here so that _sat always varies and does not lead to
// problems with rhat and neff computations
log_lik_sat[r1:r2] -= log_lik[r1:r2];
}
}
}
if (do_test) {
ppp = step((-sum(log_lik_rep) + sum(log_lik_rep_sat)) - (sum(log_lik_sat)));
} else {
ppp = 0;
}
}
} // end with a completely blank line (not even whitespace)
```
# Wrapping up
Today, we showed how to model observed data using a normal distribution
- Assumptions of Confirmatory Factor Analysis
- Not appropriate for our data
- May not be appropriate for many data sets
- We will have to keep our loading/discrimination parameters positive to ensure converges to the same posterior mode
- This will continue through the next types of data
# Next Class
- Next up, categorical distributions for observed data
- More appropriate for these data as they are discrete categorical responses
## Resources
- [Dr. Templin's slide](https://jonathantemplin.github.io/Bayesian-Psychometric-Modeling-Course-Fall2022/lectures/lecture04b/04b_Modeling_Observed_Data#/title-slide)